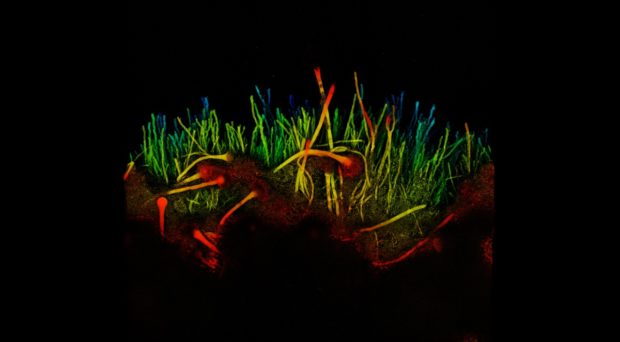
Investigating the circadian clock in Zymoseptoria tritici – Anna Tiley
One of the greatest threats facing global wheat production is the disease, “Septoria tritici Blotch”, which is caused by the fungus Zymoseptoria tritici. In Europe alone, Z. tritici is responsible for up to €700 million worth of wheat yield loss annually, and an estimated 70% of the total annual fungicide usage is targeted against it. Z. tritici can rapidly evolve resistance in the field and so novel management strategies are urgently required.
Circadian clocks are molecular machineries which are entrained by environmental signals, such as light and temperature, and which regulate key life processes. The aim of this project is to understand whether the circadian clock regulates pathogenicity and development in Z. tritici.
Z. tritici is responsible for up to €700 million worth of wheat yield loss annually, and an estimated 70% of the total annual fungicide usage is against it, with consequent risk of resistance.
Our initial results demonstrated that Z. tritici can detect light, and that this signal influences vegetative growth. We therefore hypothesised that light could be a primary input which regulates the circadian clock of the pathogen.
In the model fungus Neurosopora crassa, the circadian clock is encoded by the three genes white collar-1 (wc-1), white collar-2 (wc-2) and frequency (frq). Our bioinformatic analyses identified homologs in Z. tritici to all three of the N. crassa genes, and these were designated ztwco-1, ztwco-2 and ztfrq. Yeast-two hybrid assays demonstrated that the ZTWCO-2 and ZTWCO-2 proteins interact; a result which is also similar to observations from N. crassa.

In order to understand the role of the putative circadian clock genes in Z. tritici, we generated deletion mutants. Our results to date show that deletion of some of these candidate genes causes defects in vegetative growth, but that the mutants are still able to infect the wheat host.
This is the first in-depth study of the circadian clock in Z. tritici, and our findings open up multiple avenues for future investigation. The long-term aim of this research is to inform future control strategies against Z. tritici, such as timing of fungicide application and identification of novel targets for future fungicide screens.
 Anna Tiley is a Government of Ireland Postdoctoral Fellow based in the School of Agriculture and Food Science at University College Dublin, Ireland. She holds a degree in Biological Science from the University of Oxford, and a PhD in Molecular Plant Pathology from the University of Bristol. Anna has over seven years’ experience working with the wheat pathogen Zymoseptoria tritici, and her research focuses on the genetic basis of development and pathogenicity in this species. Follow on Twitter @tileyanna
Anna Tiley is a Government of Ireland Postdoctoral Fellow based in the School of Agriculture and Food Science at University College Dublin, Ireland. She holds a degree in Biological Science from the University of Oxford, and a PhD in Molecular Plant Pathology from the University of Bristol. Anna has over seven years’ experience working with the wheat pathogen Zymoseptoria tritici, and her research focuses on the genetic basis of development and pathogenicity in this species. Follow on Twitter @tileyanna
Exploring the variety of interactions between fungi and bacteria – Gayan Abeysinghe
I am fascinated by the way how diverse microbial species in the environment establish various modes of molecular interplay. That leads to the formation of complex microbial communities which make remarkable contributions to biogeochemical cycles, biotechnology and maintaining ecosystems.
Our group at the University of Tsukuba, steered under the guidance of Professor Norio Takeshita, contemplates the aspects of reciprocity in environmental microbial communities.
Bacterial-fungal interactions are crucial for understanding the microbial ecosystems that are closely related to agriculture, medicine and the environment.
Currently, our focus falls on fungi and bacteria which comprise a large fraction of the overall soil biomass. Bacterial-fungal interactions are crucial for the understanding of microbial ecosystems that is closely related to agriculture, medicine and the environment.
It is well known that microbial interactions are promoting the activation of cryptic biosynthetic pathways, thereby leading to the production of secondary metabolites. Those metabolites possess not only defense functions but also steer cell to cell communication and other interactive dynamics. However, the majority of studies based on the dynamics of microbiota have used monocultures.
Co-culturing has been proven to be an effective method to mimic conditions existing among microbial interactions within nature. Hence this approach may potentially facilitate the production of novel antimicrobials as well as facilitator molecules.
Together with Momoka Kuchira we were able to characterize the mutualistic relationship between the filamentous fungus Aspergillus nidulans and the gram-positive bacterium Bacillus subtilis (currently under review). Our studies provide evidence of their spatial and metabolic interaction that facilitate inter-species communication thereby exploring untraveled environmental niches and obtaining nutrients. It is understandable that there are extensive untraveled territories of microbial communities. Hence, our next milestone was to further exploit this prospect.
We conducted growth experiments of different combinations of fungal and bacterial species in coculture in selected nutrient rich and minimal conditions to observe the interactive dynamics of bacteria and fungi.
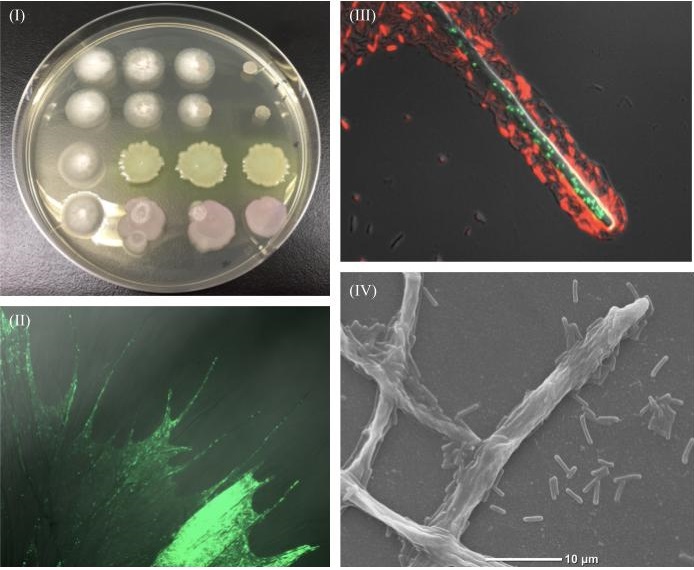
Cocultures were incubated for 1 day up to 4 days prior to microscope imaging. An array of parameters such as velocity of the movement of bacterial cells, travel distance, colonization degree and growth rate were considered to define the specificity of interaction. According to the degree of these interactions and dynamics the combinations were classified into positive, negative and neutral genres.
A selected array of combinations was subjected to LC-MS and tandem spectroscopic analysis and the differences of chemical profiles of pure and co cultures have been analyzed to determine and contrast the production levels of bioactive compounds. By chemically defining the bioactive compounds we subsequently observed the transcriptomic and genomic expression to establish an inference of their genomic potential in the state of co-existence.
We are certain that this approach would gain a more reliable perspective on the ecological context of environmental microbiota in their natural setting. Moreover, it would aid in economical and societal aspects such as bio control and therapeutic value as well as in new possible avenues for yielding antimicrobial compounds.
 Gayan Abeysinghe is a graduate student in Microbiology Research Center for Sustainability, Graduate School of Life and Environmental Sciences at the University of Tsukuba, Japan. Gayan’s research focus directs to impart insights on the microbial communication in the community setting and aims to the discovery and development of novel antimicrobials with better efficiency.
Gayan Abeysinghe is a graduate student in Microbiology Research Center for Sustainability, Graduate School of Life and Environmental Sciences at the University of Tsukuba, Japan. Gayan’s research focus directs to impart insights on the microbial communication in the community setting and aims to the discovery and development of novel antimicrobials with better efficiency.
Studying novel long noncoding RNA to understand antifungal resistance mechanisms – Danielle Weaver
lncRNA remained relatively elusive until sequencing exploded, with the discovery of genomes with small numbers of genes and surprising amounts of ‘junk’ being transcribed.
In the 1960s, DNA not coding for protein was termed ‘junk DNA’. It took another three decades before we begun to understand the gene regulatory potential of this so-called rubbish. In the 1990s, studies discovered small noncoding RNA able to silence gene translation via RNA interference. At this time, a group termed long noncoding RNA (lncRNA) were also uncovered.
Human lncRNAs such as the X chromosome-silencer, Xist, emerged first. Mechanisms included regulating gene expression by altering local epigenetics or by guiding proteins to distant genes. However, lncRNA remained relatively elusive until sequencing exploded, and we were suddenly faced with genomes with unexpectedly small numbers of genes and surprising amounts of ‘junk’ being transcribed.
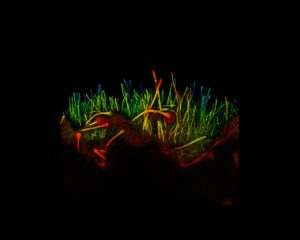
We now know of numerous regulatory lncRNA which deploy a variety of mechanisms in animals, plants and fungi. For example, Neurospora crassa produces over 2500 lncRNA. However, in the human pathogen, Aspergillus fumigatus, only a handful are known. I sought to uncover A. fumigatus lncRNA and hypothesised that they could influence its response to antifungal drugs.
Using drug exposure RNAseq experiments and a bioinformatics pipeline, we identified over 3000 lncRNA candidates. I was surprised to find over 500 lncRNA were significantly expressed upon exposure to a frontline antifungal, itraconazole. Clustering analysis showed that the lncRNA display similar dose-responsive expression patterns to genes, suggesting these lncRNA are not transcriptional noise. Instead, they are co-ordinately regulated features which may play a role in the A. fumigatus drug response.
Further investigation into these novel lncRNA may inform our understanding of antifungal drug resistance mechanisms.
 Danielle Weaver is a postdoctoral researcher at the Manchester Fungal Infection Group at the University of Manchester, UK. She completed her PhD on glycosylation in the foodborne bacterial pathogen, Campylobacter jejuni, but has since shifted focus to work on fungal pathogens. Her current research aims to harness next generation sequencing technologies to develop pathogen and drug resistance diagnostics and investigate RNA biology in Aspergillus fumigatus. Follow on Twitter @dan_weaver1 @UofMMFIG
Danielle Weaver is a postdoctoral researcher at the Manchester Fungal Infection Group at the University of Manchester, UK. She completed her PhD on glycosylation in the foodborne bacterial pathogen, Campylobacter jejuni, but has since shifted focus to work on fungal pathogens. Her current research aims to harness next generation sequencing technologies to develop pathogen and drug resistance diagnostics and investigate RNA biology in Aspergillus fumigatus. Follow on Twitter @dan_weaver1 @UofMMFIG
Synthetic control devices for gene regulation in Penicillium chrysogenum – László Mózsik
I have developed synthetic biology based-tools for scalable regulation or activation of transcriptionally silent secondary metabolite (SM) gene clusters in filamentous fungi.
Filamentous fungi produce a large variety of interesting SMs, molecules that are not essential for growth, but typically possess bioactivities that are of great value to medicine, agriculture and manufacturing. Many of these SM gene clusters are not expressed under laboratory conditions and may need to be activated or heterologously expressed before the desired products can be obtained.
Synthetic biology has revolutionized metabolic engineering and brings the exploitation of industrial microorganisms to a new level by enabling fine-tuning of gene expression allows the control of entire pathways.
Although filamentous fungi are attracting increasing interest as biotechnological production hosts, efficient genetic tools for their exploitation are limited.
Our work represents the development of synthetic gene regulatory devices, that enables scalable expression of a target gene, ranging from hardly detectable to a level similar to that of highest expressed native genes. Synthetic promoters, which were transcriptionally silent on their own, could be activated at desired level by the introduction of binding sites for the synthetic transcription factor (STF). A gene cluster may require expression levels tuned individually for each gene which is a great advantage provided by this system.
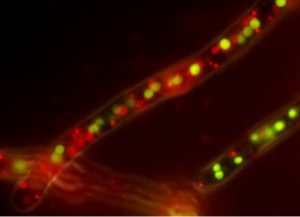
In the STF, the DNA-binding domain of the qa-1F transcription factor from Neurospora crassa is fused to the VP16 activation domain. This STF controls the expression of genes under control of a synthetic promoter containing quinic acid upstream activating sequence (QUAS) binding elements. Control devices were characterized with respect to three main features: expression of the STF, number of QUAS elements, and the type of core promoter used downstream of the QUAS element.
The versatility of the control device was demonstrated by fluorescent reporters and its application was confirmed by synthetically controlling the penicillin gene cluster in Penicillium chrysogenum for antibiotic production. We anticipate that these well-characterized and robustly performing control devices will be useful tools for silent SM gene cluster activation and for development of filamentous fungi as production hosts.
 László Mózsik is a fourth-year PhD student in the laboratory of Arnold Driessen at the University of Groningen, the Netherlands. Within the Horizon 2020 Marie Skłodowska-Curie COFUND ALERT program, his project aims at developing new genetic tools for the discovery of novel antibacterial compounds in fungi to tackle the increasing problem of multi-drug resistant bacteria.
László Mózsik is a fourth-year PhD student in the laboratory of Arnold Driessen at the University of Groningen, the Netherlands. Within the Horizon 2020 Marie Skłodowska-Curie COFUND ALERT program, his project aims at developing new genetic tools for the discovery of novel antibacterial compounds in fungi to tackle the increasing problem of multi-drug resistant bacteria.
Note: this blog post was updated on 30 June, 2020 to include an additional picture not included initially in the original post.
Anna Tiley, Gayan Abeysinghe, Danielle Weaver and László Mózsik
Gayan Abeysinghe is a graduate student in Microbiology Research Center for Sustainability, Graduate School of Life and Environmental Sciences at the University of Tsukuba, Japan.
Danielle Weaver is a postdoctoral researcher at the Manchester Fungal Infection Group at the University of Manchester, UK.
László Mózsik is a PhD student in the laboratory of Arnold Driessen at the University of Groningen, the Netherlands.
Comments