![]() A major focus of Genome Biology's RBPome issue is the role that RNA-binding proteins play in regulating splicing within the transcriptome.
A major focus of Genome Biology's RBPome issue is the role that RNA-binding proteins play in regulating splicing within the transcriptome.
But what are the triggers that cause these proteins to change their binding patterns, and so modify splicing programs? Might environmental cues such as light be responsible?
 A new article published today in our RBPome issue suggests that this might very well be the case.
A new article published today in our RBPome issue suggests that this might very well be the case.
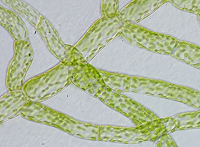
In the study, Shih-Long Tu and colleagues (Academia Sinica, Taiwan) use the moss Physcomitrella patens to report the first example of light-mediated splicing in plants.
The moss was exposed to different light conditions, following which high-throughput sequencing was employed to monitor changes in splicing. The sequencing data suggested that light induces a significant number of changes in the decisions made by protenema cells about where to splice gene transcripts.
The most common type of light-induced altered splicing event in the moss was 'intron retention'. That is, the choice of exons remained constant, but introns – which would ordinarily be excised – were in some cases retained in transcripts.
It is not clear what the consequences of intron retention are for P. patens, but one possible outcome is the production of unstable or non-functional transcripts, resulting in an overall pattern of downregulation.
Interestingly, only a subset of transcripts were targeted by intron retention, and these tended to be restricted to a small set of functions, such as 'splicing' and 'light signaling'.
This specificity not only helps support the idea that light-induced alternative splicing is a very deliberate regulatory pathway, but also points toward a mechanism for arresting certain cellular functions in response to light.
As with other forms of gene regulation, one might expect plants to be particularly expert at harnessing splicing to respond to environmental changes, given their immobility. This makes them a fascinating setting for studying the RBPome, as can also be seen in a second plant study in our RBPome issue, which examines the regulation of splicing in Arabidopsis plants subjected to salt stress conditions.
- tRFs and the Argonautes: gene silencing from antiquity - 2nd October 2014
- Keeping up with the Jobses: the role of technology in reproducible research - 26th September 2014
- How to disarm a superbug – a story told by forensic genomics - 23rd June 2014
2 Comments