The low cost computing hardware Raspberry Pi is now being used to train the next generation of computational biologists, and is proving to be a low-cost alternative to more traditional methods of learning.
“Bioinformatics is great and shouldn’t be limited to one small module” was the reaction of one enthusiastic undergraduate at the University of St Andrews (UK), following the 7 week teaching course entitled 4273 π Bioinformatics for Biologists.
They are of course correct on both counts.
Bioinformatics, or some variant of computational biology, arguably underpins a majority of modern basic biological analysis, and is working its way steadily into the realms of clinical and translational science. Think of the software you use to align your DNA sequences, infer genetic structure in your populations, or model the conformations of your newly crystallized protein. All are made possible because someone, somewhere, coded them into existence. Yet how many of us could code even a basic program of this type?
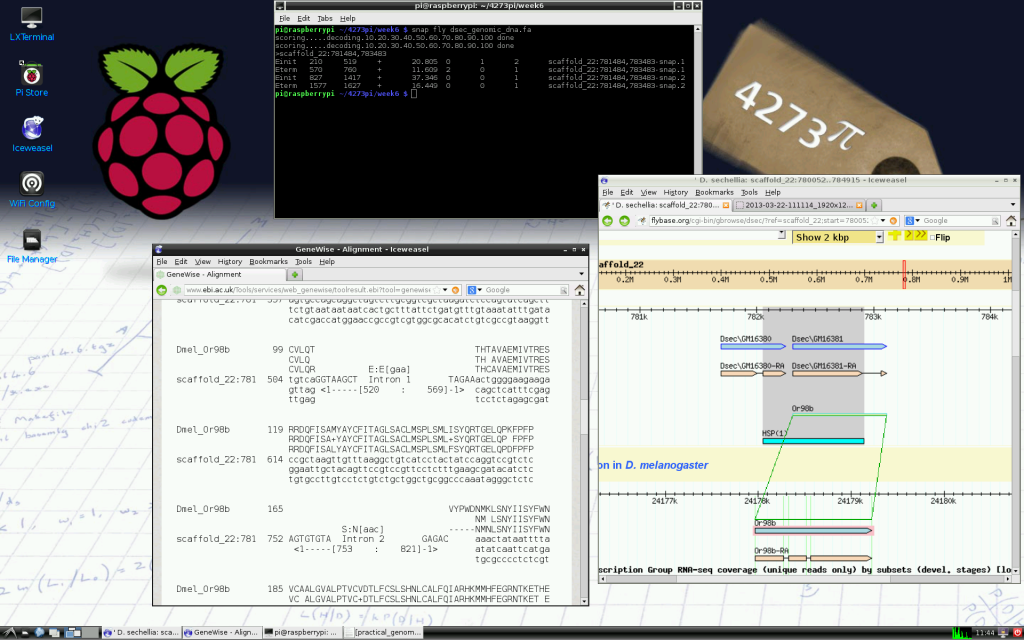
One difficulty that contributes to this issue is teaching. Few university courses exist that offer dedicated training in bioinformatics, with researchers coming to the subject either as biologists with an interest in computation, or computational scientists with an interest in biology. Although there may not be a problem with bioinformaticians coming to the field through either of these routes, training the average bench biologist or early-career researcher to have basic skills in the field can be difficult. This is partly down to the diversity of subject areas to which computational skills need to be applied.
Speaking to Biome magazine, Ian Korf, Associate Director of Bioinformatics at the Genome Center at University of California, Davis, sees teaching such diversity as a real issue for university courses “One of the greatest obstacles to teaching bioinformatics is the teachers themselves. Bioinformatics is an eclectic field drawing from molecular biology, statistics, computer science, mathematics and other disciplines. Not many teachers have such a diverse education.”
Another problem is infrastructure. Whilst some universities may have access to vast computing power for researchers, gaining access to servers that allow students to experience administrative privileges, or simply give them the time to experiment with basic computational architecture, can often be problematic.
Now, an open access, open learning method developed by Daniel Barker and colleagues aims to strip this teaching back to basics by using the newly-developed Raspberry Pi computing system to let students experience full administrator rights and gain valuable insights into real-world bioinformatics. The low-costs involved (each computer typically costs around £30/$40/€35) also means that large-scale teaching may be achieved without university costing departments having to worry about whether their laptops will be returned in full working order at the end of the semester.
What’s in the Pi?
 The hardware costs stay so low because the Raspberry Pi strips computing back to its basic elements. This credit-card sized computer eschews the modern movement toward bigger, faster processing by using a basic single-board device running an open-source operating system, without the usual hardware features like disk-drives and keyboards. Developed by a non-profit, British-based company, it is now being hailed as a revolutionary tool in facilitating mass-participation in home programming.
The hardware costs stay so low because the Raspberry Pi strips computing back to its basic elements. This credit-card sized computer eschews the modern movement toward bigger, faster processing by using a basic single-board device running an open-source operating system, without the usual hardware features like disk-drives and keyboards. Developed by a non-profit, British-based company, it is now being hailed as a revolutionary tool in facilitating mass-participation in home programming.
As well as some of the more frivolous uses to which the device can be applied, it is hoped that this low cost could not only pique a new generation’s interest in the anatomy of computing, but could also have much broader implications for access to teaching computation in the developing world. Barker feels that key to this is empowering the bench-scientist to lose their fear of the motherboard:
“Many would-be bioinformaticians get scared away because of the arcane syntax of the command line, their lack of computer programming experience, or a feeling that their mathematics skills are insufficient. These are just fears. If you hold their hand for a little while they can get through the scary bits, they will emerge on the other side self-empowered and with a new perspective on problem solving. It will impact everything from grocery shopping to genome analysis.”
A key part of this will be openness. Although developed specifically to run the bioinformatics teaching course at the University of St Andrews, Barker and colleagues acknowledge that the philosophy of openness encouraged by this new hardware also needs to be translated into teaching, and have made the course fully available to anyone wanting to have a go if they wished: full course details can be downloaded as an additional file from their article in BMC Bioinformatics.
So, if you were ever curious about what happens under the hood of your laptop, why not grab yourself a Pi and get coding?
Further reading:
Barker et al. “4273pi: Bioinformatics education on low cost ARM hardware” BMC Bioinformatics 2013, 14:243
Bioinformatics for biologists: Daniel Barker and Ian Korf discuss a novel platform for bioinformatics education. Biome magazine
Comments