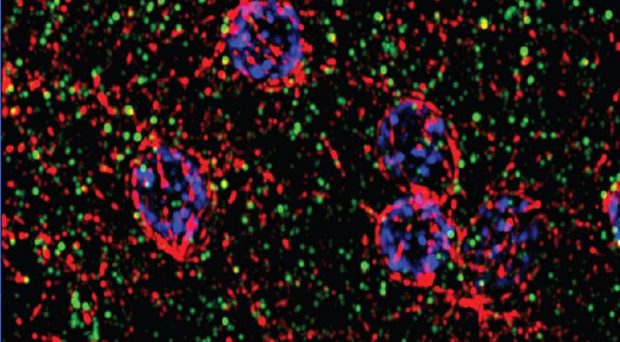
One of the most recognizable structures inside eukaryotic cells, such as human and plant cells, is the Golgi apparatus. This organelle comprises a series of membrane-bound compartments that are tightly stacked together, like a stack of pancakes. These Golgi compartments, also termed cisternae, process and sort proteins that are synthesized at the endoplasmic reticulum and destined for other organelles or the cell’s external environment.
This essential role makes the Golgi fundamentally important for healthy cellular function, and it is thought that a Golgi was present in the last common ancestor of eukaryotes, which lived approximately 1.5 billion years ago. Furthermore, disruption of the characteristic stacked structure occurs in progression of neurodegenerative diseases.
Despite the importance of Golgi stacking, how this structure is maintained is poorly understood.
Despite the importance of Golgi stacking, how this structure is maintained is poorly understood. Many different proteins have been implicated in Golgi organization in human cells. These include several rod-shaped proteins termed golgins that tether Golgi membranes to other membranes or to proteins of the cytoskeleton.
However, in some eukaryotes, such as baker’s yeast and several single-celled parasites, the Golgi is composed of compartments that are not stacked together. One such parasite is Entamoeba, which causes dysentery. The lack of stacking in such organisms raises the question of whether these eukaryotes may have lost Golgi stacking through loss of conserved genes encoding Golgi proteins.
An evolutionary approach to understanding Golgi structure
To investigate the different stacked and unstacked Golgi structures in eukaryotes, we took an evolutionary approach, as described in our recent BMC Biology article. Genomic sequence data already available from organisms across the tree of eukaryotic life allowed us to perform a comparative genomic analysis, evaluating the conservation of golgin genes across eukaryotes.
In addition, since this approach is based on comparison of eukaryotes with stacked Golgi to those with unstacked Golgi, we investigated the Golgi of a particularly relevant eukaryote for comparison: Mastigamoeba.
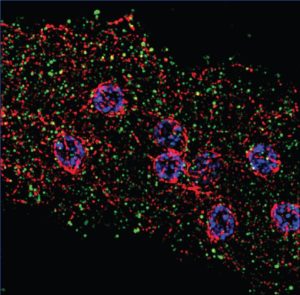
Mastigamoeba is a free-living relative of Entamoeba, and has no previously identified Golgi. However, analysis of novel sequence data from Mastigamoeba revealed the presence of genes encoding many proteins that function at the Golgi, suggesting that the Golgi must be present in some form. We then determined the structure of this hypothetical Golgi using an assay to localize the Golgi marker protein, COPI- β, which revealed unstacked compartments dispersed inside the cell (right).
An ancestral stacked Golgi apparatus
Taking these results into account, our final analysis of golgin gene conservation across a broad sampling of distantly-related eukaryotes elucidated Golgi evolution in two ways.
The highly organized stacked Golgi morphology may emerge through the cumulative activity of many different proteins, rather than one (or few) proteins acting as a linchpin
First, none of the genes we searched for tend to be lost in eukaryotes with unstacked golgi while being consistently conserved in those with stacked Golgi. This suggests, surprisingly, that the evolution of Golgi morphology cannot be explained by conservation of human Golgi stacking proteins.
Instead it seems that many diverse eukaryotes have evolved to maintain (or lose) the ancestral stacked Golgi form using different proteins than humans. This result agrees with explanations positing that the highly organized stacked Golgi morphology may emerge through the cumulative activity of many different proteins, rather than one (or few) proteins acting as a linchpin
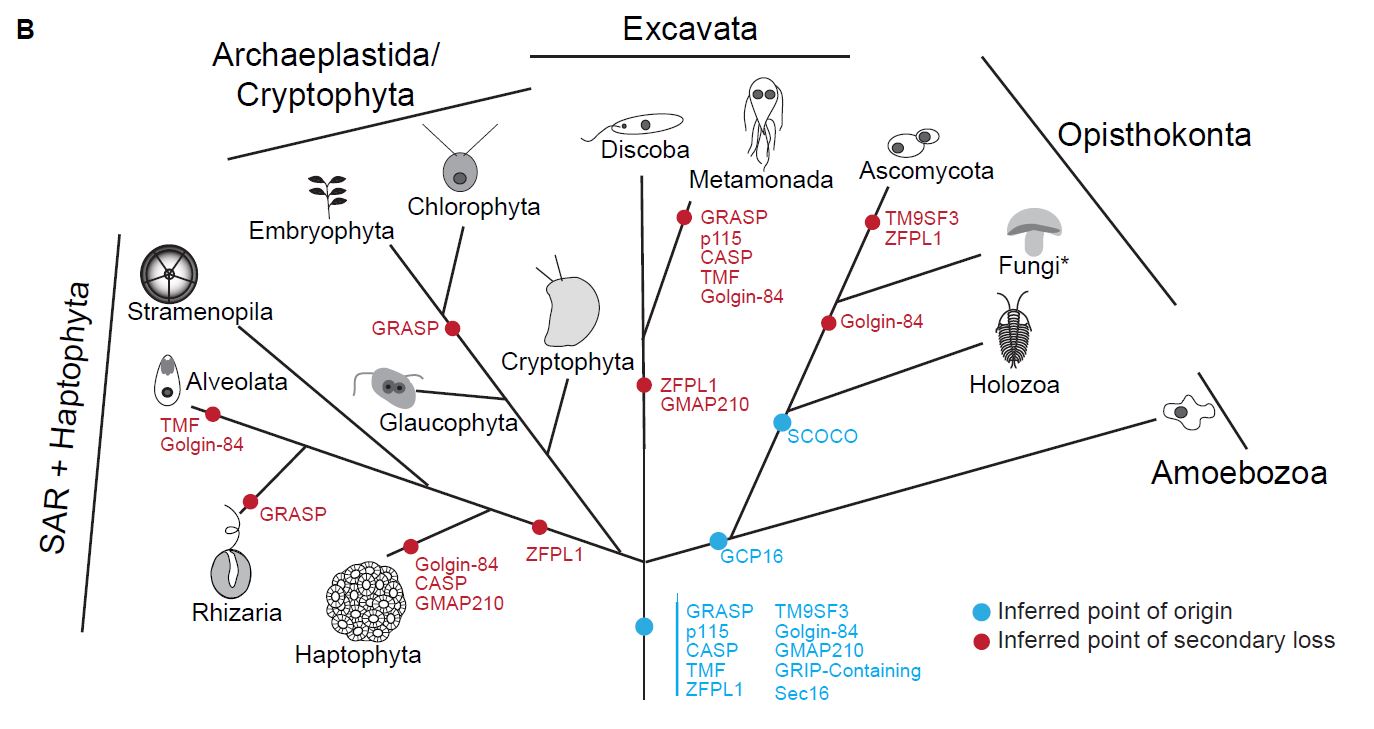
Second, although lost in some eukaryotes, conservation of some human golgins across eukaryotic diversity allowed us to infer the presence of these golgins in the last common ancestor of eukaryotes which lived approximately 1.5 billion years ago (above).
A sophisticated ancestor
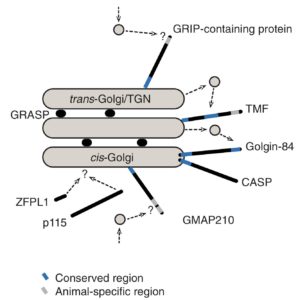
This provides a higher resolution reconstruction of the ancestral Golgi than was previously possible and suggests explicitly that it was composed of multiple functionally distinct compartments (right).
The results of this study may encourage investigation of alternative hypotheses to better understand Golgi stacking and function in both in human cells and human parasites. This study also exemplifies the utility of evolutionary approaches for constructing and testing general models of cellular processes that may be applicable to the widest diversity of eukaryotes.
The paper is published in BMC Biology as part of the “In the Light of Evolution” series.
Comments