What is your research background and why did you become interested in this?
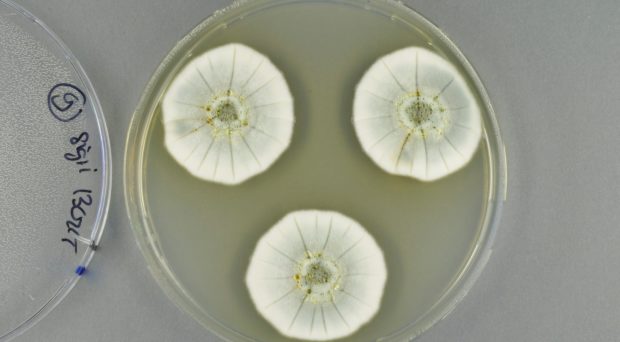
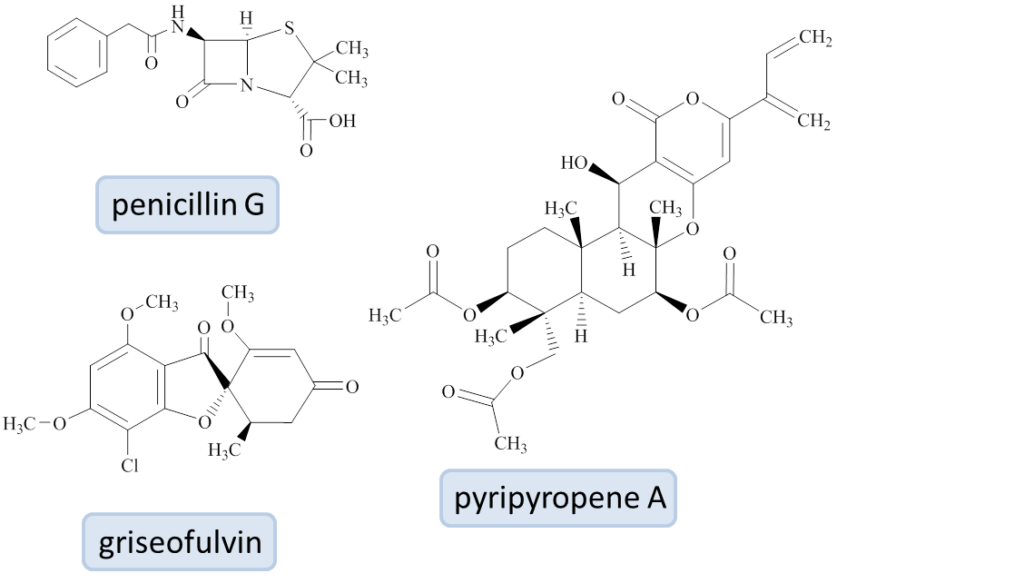
In my PhD project, I work with filamentous fungi belonging to the genus Penicillium, famous since Alexander Fleming discovered the antibiotic penicillin in 1929. Members of the Penicillium genus are also used for commercial production of other compounds, such as the cholesterol-lowering compactin and the antifungal griseofulvin.
In my project, we sequenced the genomes of ten different Penicillium species and found that they each include many genes encoding the enzymes involved in the biosynthesis of a vast array of different secondary metabolites.
These types of bio-active compounds that are produced by filamentous fungi are categorized as secondary metabolites; compounds that are not essential for growth and development of the fungus, but which often play an important role in interspecies interactions.
At the Technical University of Denmark we have a collection of more than forty thousand different fungal isolates. In my project, we sequenced the genomes of ten different Penicillium species and found that they each include many genes encoding the enzymes involved in the biosynthesis of a vast array of different secondary metabolites. As such, the genus has an enormous potential for producing secondary metabolites. This finding suggests that Penicillium species can be a source for new antibiotics and other bio-active compounds.
What was your research project about, and what did you find out?
The species in our collection come from all over the world, for example P. arizonense, that was isolated from the Grand Canyon in Arizona, USA. We found that this fungus produces a large number of interesting metabolites with antibiotic, antifungal, and anti-cancer activities such as the compound pyripyropene A.
In many cases, it is not known which genes are connected to the biosynthesis of specific secondary metabolites. This would be valuable information if, for example, you like to optimize the production of a given compound. The sequencing of different fungal genomes can help in establishing these links; you can find two fungi that both make the same secondary metabolite, and look for similarities in their genomes.
What brought you to work with fungi?
During my studies at the Technical University of Delft, I learned to see micro-organisms as (cell) factories that can produce all kinds of chemicals for us. In Delft I mainly worked with yeast as a cell factory, but when I learned more about filamentous fungi and their great capacity to produce enzymes, organic acids and secondary metabolites I was intrigued. Now I am part of the Biosynthetic Pathway Engineering group at the Technical University of Denmark.
Where would you like the research to take you?
I am really excited about the fast development in the field of industrial microbiology and, after I finish my PhD project at the end of this year, I would like to continue studying the huge potential of fungal cell factories for a more bio-based future.
What advice would you give to young scientists starting out in their career?
My PhD project is part of a bigger Marie Curie Initial Training Network, called QuantFung.
I recommend everyone to take part in such training networks. . It is a great way to meet researchers in an international network, to start collaborations, and to make friends.
And one more thing: be happy with all successes, no matter how small they are. And most importantly, celebrate them! I loved that my supervisor already got champagne after submission of our manuscript, instead of after acceptance!
Enjoyed reading this blog? Read our other Fungal Genetics Conference Q&As here:
Comments