A series of articles in the BMC series and GigaScience accompany the publication of the swine genome sequence
Today sees the publication of the genome sequence of the domestic pig in Nature by the Swine Genome Sequencing Consortium and sequencing of the Wuzhishan miniature pig in GigaScience . Accompanying these publications are a litter of companion articles in a BioMed Central cross-journal article series.
Yutao Du, Shutang Feng and colleagues describe the independent genome sequencing of the Wuzhishan miniature pig in the journal GigaScience. This breed is an up and coming model for human medical applications and its sequencing provides important information about genetic similarities and differences between the genes involved in coronary artery disease and drug targets in humans and pigs. Also they found that the breed has lost one species of virus incorporated into the genome, unlike other pig breeds, increasing the potential for xenotransplantation between the pigs and humans.
 |
 |
Duroc (left) and Wuzhishan (right) pigs: subjects of the two sequencing projects. Credit: Duroc adapted from image by David Merrett on Flickr, Creative Commons 2.0
Medical Models
Several articles in the series explore the biomedical importance of the genome sequence. Eric Walters and colleagues review the role of the pig as a large animal biomedical model and how the pig genome sequence improves our understanding. They explain that the pig is a very good model of cardiovascular disease, and the only animal to accurately recapitulate cystic fibrosis symptoms.
Tom Freeman and colleagues describe the first detailed gene expression atlas of the domestic pig. This provides an important new resource for study of the physiology of mammalian tissues. They demonstrate this with a detailed look at the gastrointestinal tract, highlighting several candidate genes for gastrointestinal diseases. Other articles in the series survey prohormone and convertase genes, the olfactory gene repertoire and beta-defensin genes in the pig genome.
Meaty findings
Other articles focus on the relevance of the genome sequence to understanding the pig as food animal. Pamela Wiener, Rob Ogden and colleagues describe a new genetic diagnostic assay able to distinguish between British traditional pig breeds. The authors explain that mislabelling is prevalent in the food industry but their assay can successfully distinguish pork products from different traditional breeds that often command a price premium over commercial breeds.
Finally, some further articles explore the structure of the pig genome, and the assembly and annotation of its sequence. In one article Martien Groenen and colleagues mapped recombination across the genome, finding sex-specific recombination rates and correlations with GC content. Bertrand Servin and colleagues describe the use of radiation hybrid mapping in assembly of the genome.
All these articles and more are available to read on the series homepage.
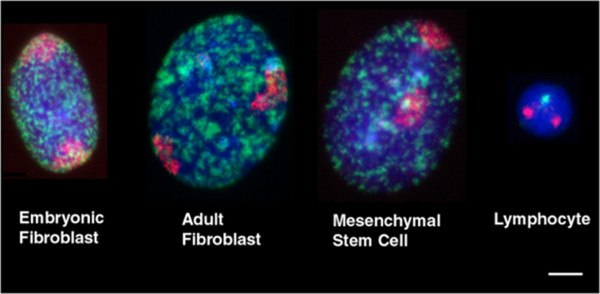
Images of porcine interphase chromosome territories in nuclei derived from embryonic and adult fibroblasts, mesenchymal stem cells and lymphocytes. Taken from Foster et al. BMC Cell Biology 13:30
Tim Sands
Latest posts by Tim Sands (see all)
- Companion articles to the swine genome sequence - 15th November 2012
Comments