Re-evaluation of the Haarlem Archaeopteryx and the radiation of maniraptoran theropod dinosaurs

It is 1861 and Hermann von Meyer, a German paleontologist, inspects a slab of Bavarian limestone depicting a single feather. He will name this specimen archaeopteryx, signifying an “ancient wing” or “feather”.
The same year, the first Archaeopteryx skeleton was discovered, depicting what was to be considered the oldest known ancestor of the modern bird for decades to come. Marking the boundary between bird and dinosaur, this iconic fossil has been pivotal to our understanding of avian evolution.
In this study, the authors re-examined one of the least complete specimens of Archaeopteryx, currently located in the Teylers Museum in Haarlem. After conducting anatomical analysis, they concluded the “Haarlem specimen” was more likely to have descended from the Chinese Anchiornis, a group of winged dinosaurs that predate the Archaeopteryx.
This study has had fascinating implications for our understanding of the origins of modern birds. The authors propose a westward spread of the avian ancestors from their origins in East Asia and arriving in Bavaria, where their remains would be found 350 million years later.
A new species of Xenoturbella from the western Pacific Ocean and the evolution of Xenoturbella

From the extinct to the extant, we move on to the discovery of a new species of Xenoturbella; a truly bizarre marine organism. Until recently, this mysterious creature had only been collected in the seas off the coast of Sweden.
It would not be difficult to mistake Xenoturbella for an old sock, lying abandoned on the sea bed. Its morphology is very simple and its behavioral habits unknown. Even its phylogenetic position has been keenly debated throughout recent years.
Perhaps Xenoturbella’s most defining characteristics are the absence of an anus and the lack of a centralized nervous system. Its mouth serves as the single opening through which food and waste can be exchanged. These remarkable traits make it an attractive choice for investigating the evolutionary origins of these features.
A paper published in BMC Evolutionary Biology this year reports the identification of a new species of Xenoturbella in the Western Pacific Ocean off the coast of Japan. It is relatively easily collected via marine biology dredge and thus holds a lot of promise as a research organism for the further characterization of this strange species.
Epigenetic variation between urban and rural populations of Darwin’s finches
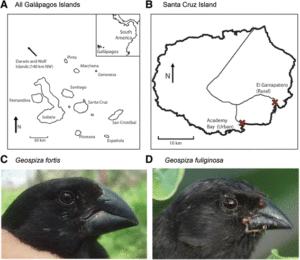
Travelling across to the far side of the Pacific Ocean, the authors of our next paper go to the Galapagos Islands to study Darwin’s finches. These birds are best known for helping shape Charles Darwin’s theory of natural selection. There are 16 species native to the Galapagos islands, each most clearly identified by their differing beak sizes and shapes that are adapted to the different food sources found on each island.
Nearly 160 years later, the field of epigenetics may provide an explanation for the rapid adaptation that these finches display. Epigenetics refers to the study of changes in gene expression rather than the genetic code, achieved through modifications of DNA by processes such as methylation.
DNA methylation describes the addition of a carbon and three hydrogen atoms to a cytosine nucleotide, when followed by a guanine, in the order “CpG”. Some methylation patterns have been shown to be heritable, while methylation patterns have even displayed a mutation rate higher than that observed in the genetic code. Crucially, for this study, the environment has also been reported to influence DNA methylation patterns.
This points to the possibility that epigenetics may contribute to quick phenotypic changes within each species of Darwin’s Finches. However, the process through which this could happen is still unknown.
After studying two species of finch, the authors reported significant differences in methylation patterns between urban and rural populations of each species. Urbanization in the Galapagos Islands has happened within the last 60 years, so while the authors are careful to acknowledge the possibility of random epigenetic drift, if urbanization is the cause for these differences in methylation then these changes have occurred impressively quickly.
A genetic chronology for the Indian Subcontinent points to heavily sex-biased dispersals
 Our next study examines mitochondrial DNA to determine the ancient ancestry of the modern day Indian population. Between 200,000 and 400,000 years ago, anatomically modern humans were thought to have moved out of east Africa and through the Indian subcontinent, consequently making the Indian population one of the most genetically diverse on the planet.
Our next study examines mitochondrial DNA to determine the ancient ancestry of the modern day Indian population. Between 200,000 and 400,000 years ago, anatomically modern humans were thought to have moved out of east Africa and through the Indian subcontinent, consequently making the Indian population one of the most genetically diverse on the planet.
However, the genetic study of the Indian population is no easy task. High levels of endogamy are common due to rigid social boundaries imposed by the caste system. The upshot of this is variation in the genetic code, known as genetic drift, between these socially isolated populations.
Perhaps most interestingly, a movement of people from central Asia around 4000 years ago provides new evidence for the spread of the ancient Indo–European language into India.
The authors of this study aimed to remedy these problems by tracing the inheritance of mitochondrial DNA, which is exclusively maternally inherited as well as DNA found exclusively on the Y chromosome, which is exclusively passed down the male line. This allows lineages to be traced for millennia back in time. In combination with genome wide analyses, ancient immigration patterns were able to be deduced.
The theory proposing an “Indo-Aryan invasion” to India is highly controversial. It has the potential to explain the presence of Indo-European languages, spoken across the northern half of India, while some have also attributed the origins of the caste system to this hypothetical event. Previously, there had been no archaeological data or genetic data to support any claims this event may have occurred.
What is proposed in this paper is a succession of migration waves from Iran, the Caucasus and Anatolia over the period of 50,000 years. Perhaps most interestingly, a movement of people from central Asia around 4000 years ago provides new evidence for the spread of the ancient Indo–European language into India.
Skeptics have pointed to the lack of ancient preserved DNA in South Asia to confirm any migration occurred. This study does, however, provide an exciting new perspective on this often disputed topic.
Going deeper in the automated identification of Herbarium specimens

The final study in our end of year selection focuses on the use of artificial intelligence to identify new species of plant. Vast numbers of preserved plant specimens have been increasingly archived in a digital format and placed online in publicly available databases.
As contemporary understanding of plant phylogenetics and evolution develops, our knowledge of plant species becomes outdated and data entries must be revised. Around 350 million specimens are stored in herbaria around the world. This huge amount of data means that this is no fitting task for mere mortals.
This year, Carranza-Rojas et al designed an algorithm designed to use deep learning to identify plant species based on digital images of preserved plant specimens. The software was able to produce the correct plant species within the top 5 results 90% of the time, while the correct result was the first hit for 70% of cases.
Deep learning technology shows considerable promise in the field of botany, with the potential to revolutionize plant identification. We look forward to further studies and new break-throughs, ushering in the next generation of software in the digital age.
Comments