
Photosynthesis is a complex pathway in which light energy and water are used to fix carbon, in the form of carbon dioxide, into sugar with an oxygen by-product. Plants, algae, and cyanobacteria rely on this process to produce their own source of energy, while we benefit from the creation and maintenance of both the Earth’s oxygen content and the primary source of food.
There are three types of photosynthesis, defined by the carbon fixation pathways used: C3, C4 and CAM.
Plants with C4 photosynthesis have evolved independently 45 different times throughout the history of flowering plants. However, only about 7600 plant species, or 3% of all known plant species, use this pathway. Evolution of the 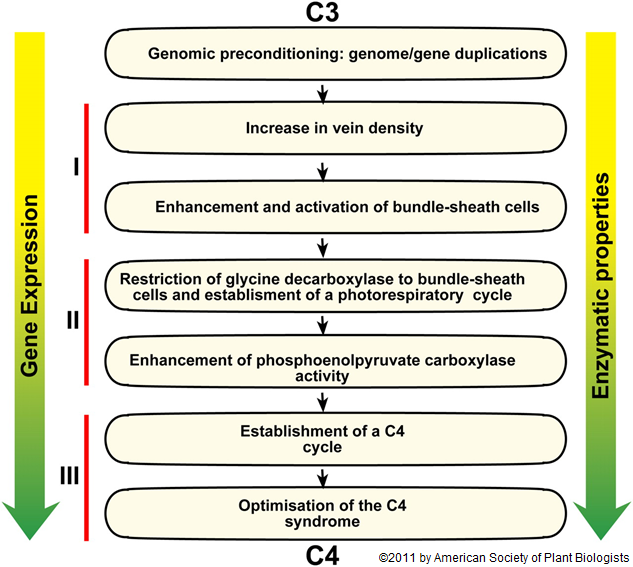 C4 pathway involves alterations in leaf anatomy, gene expression, cell biology, and biochemistry.
C4 pathway involves alterations in leaf anatomy, gene expression, cell biology, and biochemistry.
But C4 plants have a competitive edge over C3 plants: they use 3-fold less water, allowing them to grow in conditions of drought, high temperature, and carbon dioxide limitation.
Many researchers are now focused on understanding the differences between C3 and C4 plants, in an attempt to engineer important crops to be more energy and water efficient. Only four major crops use C4 photosynthesis: maize, sugar cane, sorghum, and millet.
However, there are bioengineering projects underway with the aim of increasing productivity in food and biofuels. To understand the evolution of C4 metabolism and how it can be engineered, we need to have genomic and transcriptomic information from closely related C3 and C4 species.
Comparisons between C3 and C4 plants have mostly focused on the known C4 crops and important C3 crops like rice and wheat. But the two groups are not very closely related, having diverged 50 million years ago. Distant evolutionary relationships can sometimes fail to detect the genomic changes associated with the evolution of C4 photosynthesis.
A recent study published in Genome Biology by Anthony Studer and colleagues at the Donald Danforth Plant Science Center helps to bridge this gap by presenting the genome and transcriptome of Dichanthelium oligosanthes. This species of panicoid grass diverged from maize, sorghum, and sugarcane 15 million years ago, before the evolution of C4 in that clade. In other words, D. oligosanthes is a C3 panicoid grass that is closely related to important C4 panicoid grass crops.
The authors present a draft quality assembly of the genome of D. oligosanthes, representing 78% of the estimated total genome, but covering 98% of the coding sequence. They then looked at gene expression profiles of D. oligosanthes and a closely related species of millet over different stages of leaf development, as well as the expression patterns of the six core C4 enzymes. This gave insights into changes in gene regulation linked to the evolution of the C4 pathway.
Studer and his colleagues hope that the genomic information they have provided on D. oligosanthes, the only C3 grass in a C4-rich clade, will facilitate the study of the mechanisms involved in the evolution of C4 photosynthesis. They also hope D. oligosanthes can be used as a model for studying cold tolerance in panicoid grasses to produce heartier crops for growth in the Northern hemisphere.
Dominique Morneau
Latest posts by Dominique Morneau (see all)
- Best of 2017: Top Picks from Genome Biology - 15th December 2017
- Exploring the epigenetic dynamics of early plant development - 25th September 2017
- Fascinating Plant Epigenomes - 18th May 2017
Comments