
Evolutionary Genomics
 Researchers in South Korea assembled the genomes of indigenous and domesticated African cattle populations, unraveling their unique adaptive diversity and providing clues to promote sustainable livestock improvement.
Researchers in South Korea assembled the genomes of indigenous and domesticated African cattle populations, unraveling their unique adaptive diversity and providing clues to promote sustainable livestock improvement.
A look at genetic variation in Indian and Pakistani Parsi populations, as well as ancient Parsi DNA samples, revealed that Parsis are more genetically similar to Neolithic Iranians than modern Iranians, who have experienced a recent wave of admixture from the Near East.
There is 1.5-4% Neandertal ancestry in present-day non-African humans. We learned that the major influence of Neandertal introgressed alleles is on gene regulation.
Crop Genomics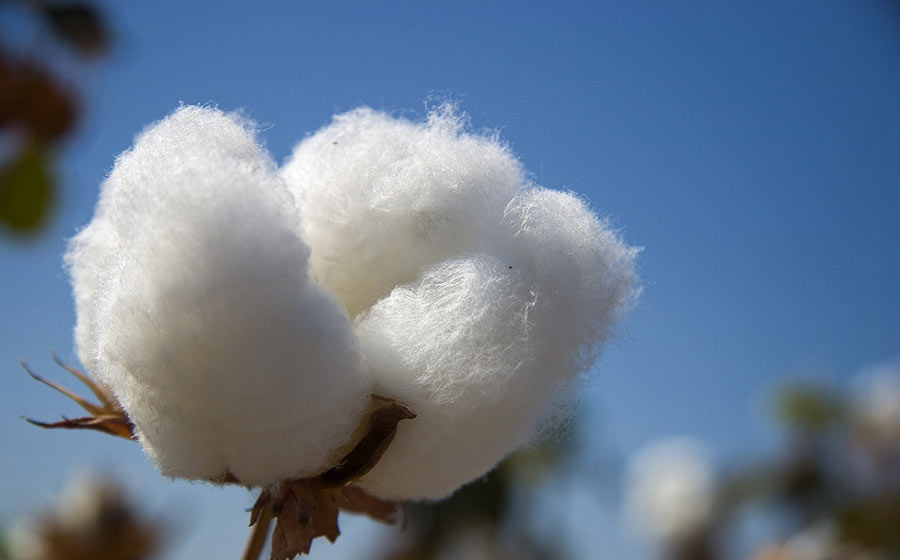
Population analysis of 29 accessions of common bean gave us evidence an unprecedented speciation event in the tropical Andes, giving rise to a sister species, which was previously thought to be the wild ancestor of Phaseolus.
We also got some insights into the domestication of two species of commercial cotton.
Cancer and Disease Genomics
A novel primate-specific lncRNA was found to be a contributor to the complex mechanisms underlying the development and progression of colorectal cancer, and could potentially play a role as a biomarker for the disease.
Researchers from the Baylor College of Medicine compared the performance of 25 algorithms for variant classification, providing combinations of tools that can be used in the assessment of clinically relevant variants in accordance with the ACMG/AMP guidelines.
Whole-exome sequencing analysis identified a number of candidate genes potentially associated with known Parkinson’s Disease mechanisms. Functional analysis of these genes in Drosophila provided additional evidence of their function in disease progression.
We learned about the structural variants in the human genome associated with disease, and that the diversity and abundance of complex structural variants in the morbid human genome is far larger than previously anticipated.
Epigenomics
Potential new avenues for inducing epigenetic and genetic diversity for crop breeding were open up by the finding that inhibition of RNA Polymerase II leads to a burst of transposon activity in Arabidopsis and rice, providing.
 Researchers at the University of Texas at Austin examined the epigenomic signatures of domestication traits during cotton evolution, which may have contributed to the crop’s worldwide cultivation.
Researchers at the University of Texas at Austin examined the epigenomic signatures of domestication traits during cotton evolution, which may have contributed to the crop’s worldwide cultivation.
In the world of mammalian epigenomes, were given the mouse epigenetic clock, which will be invaluable for understanding the biology of aging and will provide avenues for manipulating (and possibly resetting) its speed in vivo.
Technological Advances
Using the diagnostic potential of cell-free DNA, researchers from UCLA created a probabilistic method for not only detecting tumors, but also their location. CancerLocator has been accessed over 19,000 times so far and created quite a buzz in the news!
We got a step closer to unlocking the full potential of metagenomic data with DESMAN, which identifies variants in core genes and links them into haplotypes and abundance profiles, allowing species-level identification.
 Easi-CRISPR, a new method for animal genome editing, addresses the problem of inefficiency of targeted DNA casette insertion. In the development of the method, the authors produced correctly targeted alleles in 100% of live offspring.
Easi-CRISPR, a new method for animal genome editing, addresses the problem of inefficiency of targeted DNA casette insertion. In the development of the method, the authors produced correctly targeted alleles in 100% of live offspring.
We also saw the first application of the CRISPR/Cas9 system in cattle. Researchers from China targeted the NRAMP1 gene with reduced off-target effects, creating transgenic cattle with increased resistance to tuberculosis. This is among our most talked-about articles of the year!
Single-cell analysis got a bit of lasting power with a method for sample preservation that maintains transcripts in viable single cells, disconnecting the logistics of sampling from the subsequent processing and analysis steps.
Microbial Genomics
 A microbiome profile of 531 Finnish men identified links between gut microbiome composition and circulating metabolites, teasing apart more of the story in the human-microbiome relationship.
A microbiome profile of 531 Finnish men identified links between gut microbiome composition and circulating metabolites, teasing apart more of the story in the human-microbiome relationship.
The first methicillin-resistant Staphylococcus aureus isolates, preserved since the 1960s, showed us that antibiotic resistance can predate the widespread use of an antibiotic, demonstrating that new drugs could be rendered ineffective by previous adaptations to bacterial genomes. Listen to Scientific American cover the article in their 60-second Science podcast!
Thank you to all our authors and reviewers in 2017. We look forward to a lot of great research to come in 2018!
Dominique Morneau
Latest posts by Dominique Morneau (see all)
- Best of 2017: Top Picks from Genome Biology - 15th December 2017
- Exploring the epigenetic dynamics of early plant development - 25th September 2017
- Fascinating Plant Epigenomes - 18th May 2017
Comments