Darwin’s ideas about cattle genetic origins and domestication
Charles Darwin was fascinated by the process of domestication and made an excellent case that subjugation and artificial selection of animals and plants echoed – albeit at a more rapid pace – evolution via natural selection in the natural world.
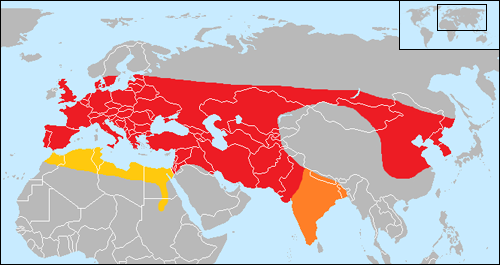
During the course of marshalling his evidence for biological evolution, Darwin also contemplated the origins of domestic cattle, realizing correctly that humpless taurine and humped zebu cattle had been developed by humans from biologically distinct populations of the now-extinct wild aurochs (Bos primigenius) that ranged across Eurasia during the last days of the Ice Age.
“In regard to sheep and goats I can form no opinion. I should think, from facts communicated to me by Mr. Blyth, on the habits, voice, and constitution, &c., of the humped Indian cattle, that these had descended from a different aboriginal stock from our European cattle; and several competent judges believe that these latter have had more than one wild parent.”
Chapter 1: Variation under Domestication, On the Origin of Species (1859)
This quotation from On the Origin of Species shows that 19th century cattle breeding experts considered it likely that European cattle had multifaceted genetic origins with “…more than one wild parent.”
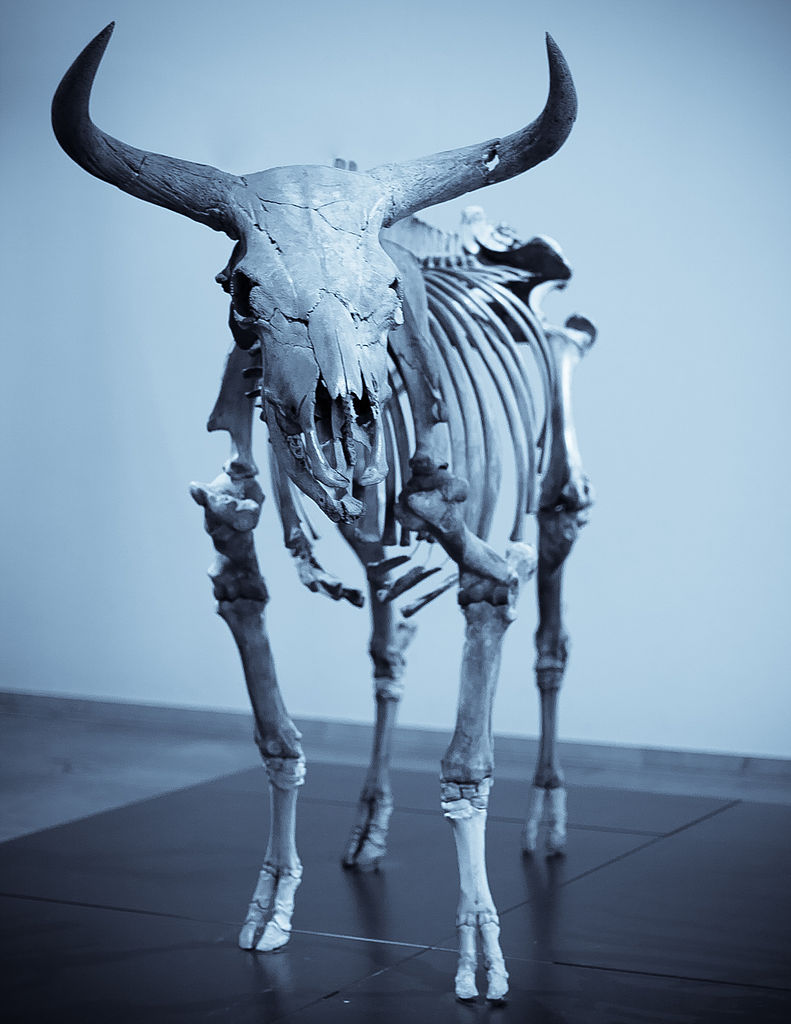

With the publication of the first complete nuclear genome sequence from the wild aurochs, it is now clear that these Victorian breeders were remarkably prescient. Simple models of domestic cattle evolution based on analyses of uniparental mitochondrial and Y chromosome DNA, which we and others proposed, need to be revised towards a more nuanced picture of crossbreeding and gene flow between domestic cattle and wild aurochs, as early European farmers moved into new habitats, such as Britain during the Neolithic.
What did we do?
In 2001, we published a survey of mitochondrial DNA variation in domestic cattle and a number of British aurochs bone specimens, concluding that wild aurochs made little or no contribution to the gene pool of modern European cattle.
Ten years later, in 2011, high-throughput next-generation sequencing technology had begun to sweep all before it, including reinvigorating the study of ancient DNA and giving rise to the new field of palaeogenomics.
This encouraged us to start a genome sequencing project using DNA extracted from an aurochs humerus bone originally excavated in 1998 from a Derbyshire cave, which displayed remarkable DNA preservation even though it was almost 7,000 years old.
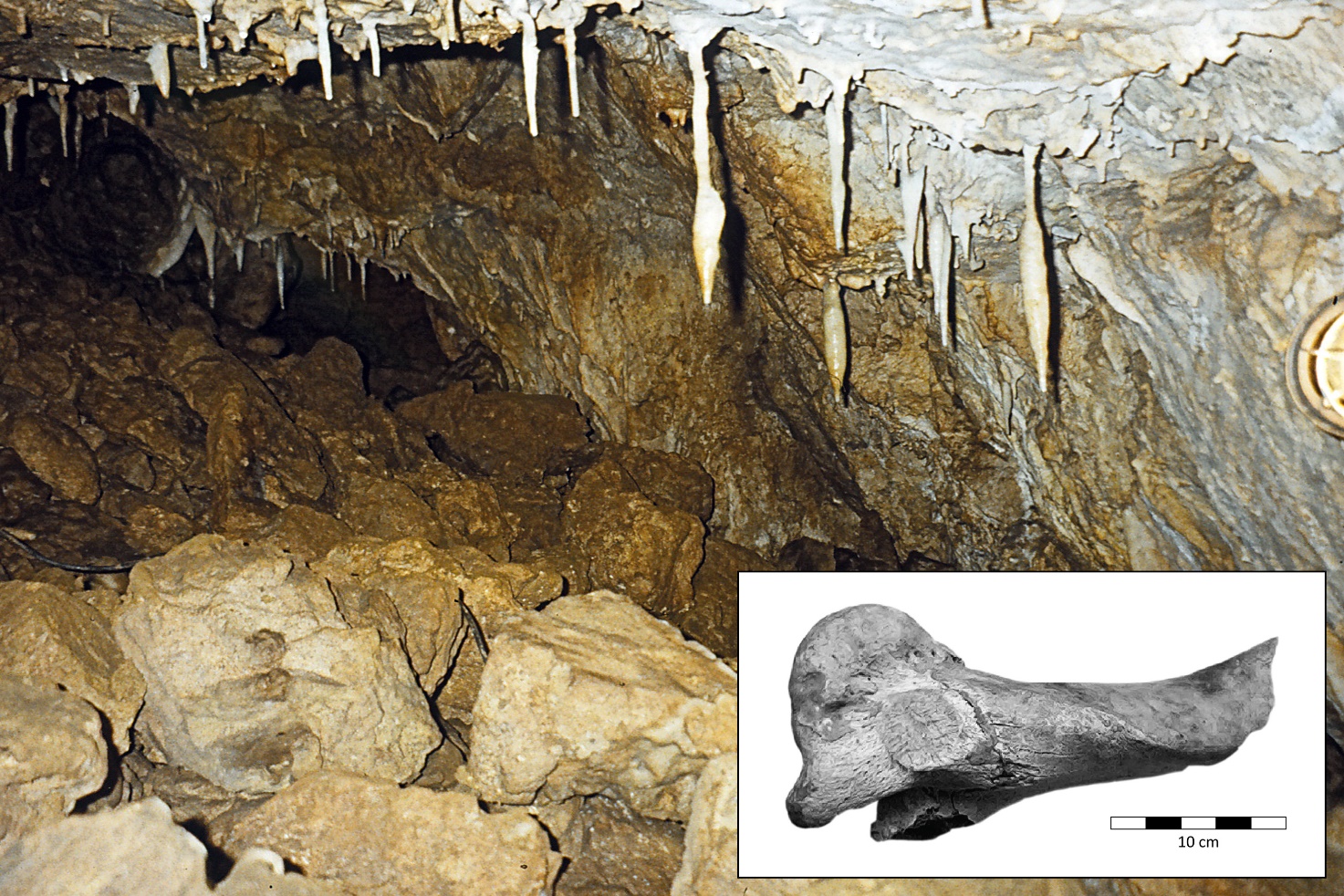

Using ancient DNA laboratory facilities at the Molecular Population Genetics Lab at Trinity College, Dublin, we systematically generated multiple genomic libraries of DNA fragments from the Derbyshire aurochs bone.
These were then used for high-throughput DNA sequencing to obtain 3.37 billion DNA sequence reads, 417 million (12.4%) of which could be aligned with high confidence to the cattle reference genome sequence.
Lead author Stephen Park then analyzed this aligned aurochs genome sequence in conjunction with whole genome sequence data from 81 modern domestic cattle samples (both humpless B. taurus and humped B. indicus), which was generated by our co-authors at the USDA Animal Genomics and Improvement Laboratory.
We were also able to compare the aurochs genome sequence to high-density single nucleotide polymorphism (SNP) genetic marker data from more than 1,200 cattle representing a wide variety of taurine and zebu breeds from across the globe.
What did we discover?
Once we had generated our Derbyshire aurochs genome sequence, we constructed phylogenetic trees using genetic marker data from modern cattle.
These evolutionary trees suggested that the population of aurochs present in Britain prior to the arrival of agriculture to the island – approximately 5,500 years ago – were genetically distinct from the aurochs populations domesticated by early farmers in the Middle East more than 4,000 years previously.
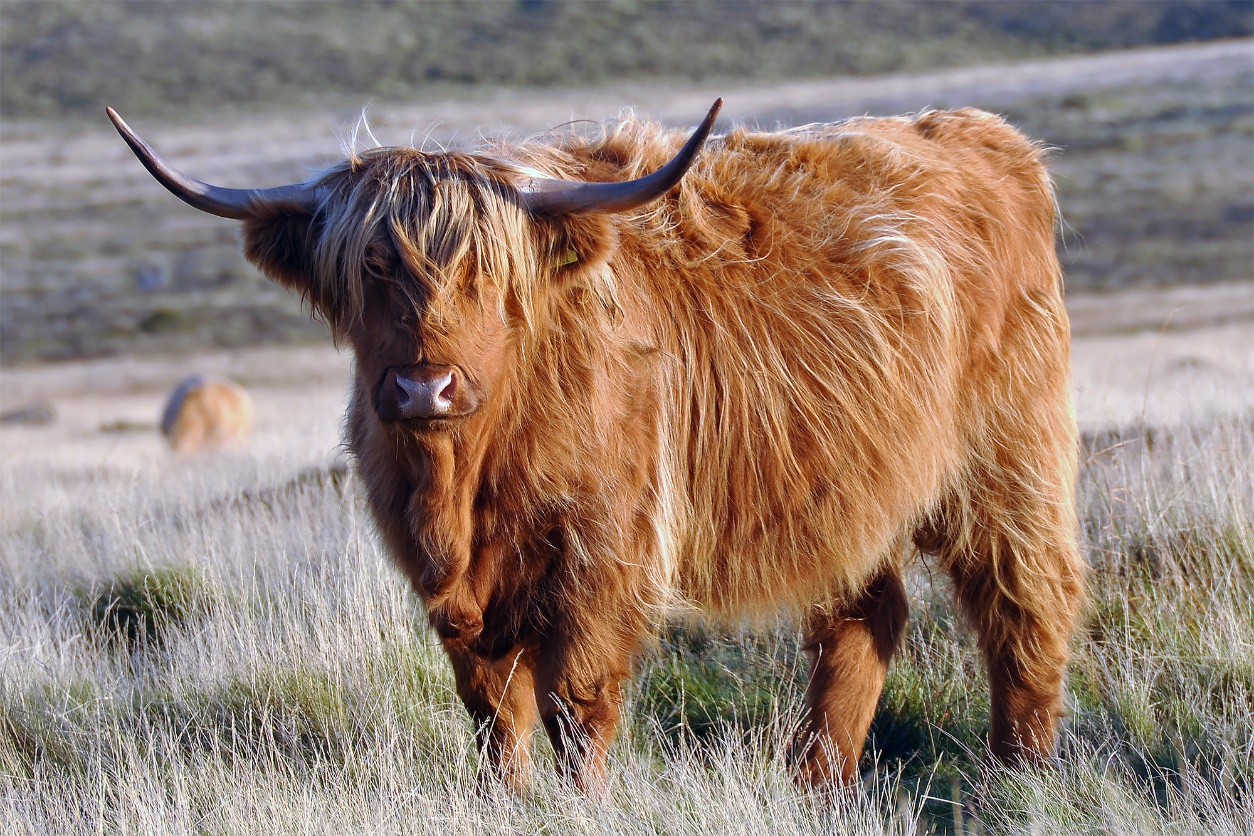
To our surprise, however, higher-resolution analyses of the Derbyshire aurochs genome with data from modern European cattle revealed an ancient legacy of crossbreeding and gene flow between wild aurochs and the ancestors of modern British and Irish cattle, which was particularly evident for heritage or landrace breeds such as Scottish Highland and Irish Kerry.

The second phase of our research work used comparative analyses of the Derbyshire aurochs genome with genome sequence information from 81 modern cattle to determine how domestication has shaped the genomes of modern taurine cattle.
These analyses implicated genes involved in brain development, muscle growth, metabolism and the immune system and resistance to infectious disease. These observations tallied well with previous work in other domestic animals, including dogs, cats, pigs and rabbits, suggesting that domestication and early human-mediated selection has targeted similar sets of genes in different mammalian species.
What does it mean for cattle agriculture today?
This first aurochs genome sequence will provide an important comparative reference to help animal geneticists and breeders build a clearer picture of the genetics underlying important behavioural, production and health traits in modern domestic cattle populations.
This information will be particularly valuable for the genome-assisted breeding programmes that increasingly underpin dairy and beef cattle production in many countries.
Notwithstanding the outputs relevant for commercial cattle breeding, the aurochs genome project has revealed that British and Irish ancient heritage cattle breeds can trace a significant portion of their genetic make-up back to the magnificent wild aurochs that roamed Britain prior to the appearance of farmers on the island.
These results will therefore have significant implications for genetic conservation of unique breeds such as Scottish Highland and Irish Kerry cattle.
David MacHugh
He received a B.A. (Mod. Genetics) honours degree from Trinity College Dublin and did a PhD in molecular population genetics also at Trinity College Dublin in 1996.
His current research focuses on population genomics and functional genomics of performance and health traits in domestic livestock.
Latest posts by David MacHugh (see all)
- The gruelling hunt for the aurochs genome - 27th October 2015
Comments