In 2014 we are awash with data. Resequencing is modestly priced and a range of platform technologies provide terabytes of “-omics” information about the world’s great infectious diseases. On the parasites that cause them, the vectors that transmit them and on the host response to them. It seems a long time now since the the Tritryp (Trypanosoma brucei, Trypanosoma cruzi and Leishmania major) genomes were first analysed and published. In the intervening period a wealth of attendant data from other platforms have provided gene expression data for life-cycle stages via the proteomes and transcriptomes, and new strains of these parasites are being sequenced every day, providing us with a detailed knowledge of the heterogeneity present in the gene pool of each of these species.
Surprisingly, one of the four human infective trypanosomatid species had apparently slipped out from under the gaze of the early sequencing networks and consortiums. A trypanosome which infects a very large number of mammals and animals in Latin America and which is vector borne – transmitted with, alongside of and by the same vectors that transmit Chagas disease, but which never causes disease and which has a considerable cost to the Latin American health systems by confounding diagnosis and confusing treatment. The parasite is “the other American Trypanosome” Trypanosoma rangeli.
The significance of T. rangeli is not though confined to confounding treatment and diagnosis of Chagas disease. Along with its interesting and distinct biology, the high degree of shared antigenicity may make it actually protective against Chagas disease, while it’s pathogenicity to triatomine vectors makes it a potential biological control agent. Perhaps most importantly, its avirulence and recent divergence from T. cruzi make it an ideal candidate for comparative genomics and drug/vaccine candidate screening. Especially, when compared with genomes from each of the major pathogenic lineages of T. cruzi and an apparently asymptomatic lineage associated with bats – T. cruzi marinkellei.
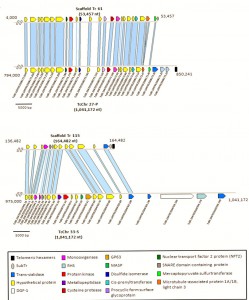
This week has seen the publication of the first T. rangeli genome and its deposition in TriTrypDB/EuPathDB by a consortium orchestrated from UFSC (Florianópolis) in Brazil. As might be expected most (>90%) of its genes are shared with T. cruzi, but it has far fewer members of the multicopy virulence factor families such as trans-sialidases, mucins and MASPs and conversely more copies of the amastin family of genes suggesting a role for them in vector interaction. Interestingly evidence of the RNAi pathway was found and although it appears to be non-functional it is indicative of the progressive loss of the pathway in this lineage of trypanosomes. Finally, the antioxidant defenses are noticeably deficient which may relate to poor intracellular survival and the lack of virulence in the mammalian host.
The key to Chagas virulence will lie in the differences between the avirulent and the virulent as it does for other parasites where avirulent strains and relatives are increasingly the focus of new research. It is to be hoped that the publication of this genome and the genomes of other related trypanosomes will open the door for the comparative analyses which will provide a comprehensive view of Chagas disease pathogenesis and perhaps point the way to the most effective new lines of treatment which remain sorely needed.

Comments