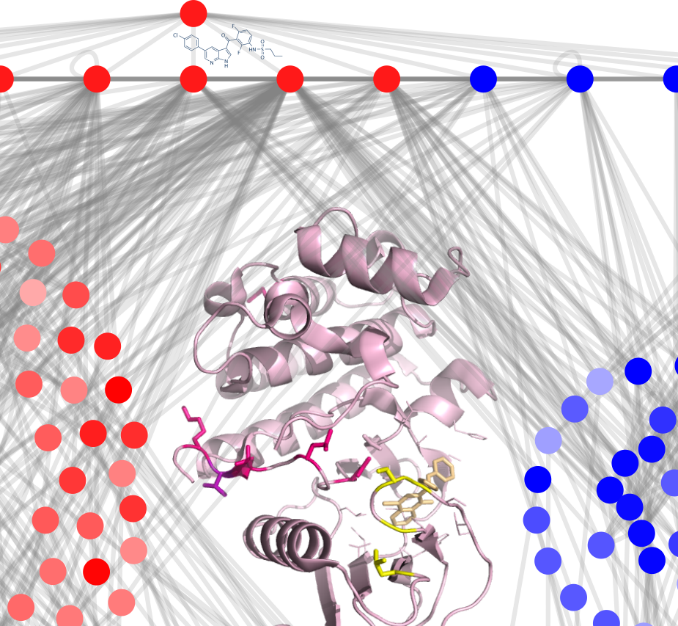
A hidden layer of regulation selects surviving cancer cells
It is widely established that cancers develop when cells mutate. However, resistance to therapy might go beyond cancer mutations. Though chemotherapy might kill most of the cancer, tiny populations of drug-resistant cancer cells manage to survive and propagate. Unlike the more familiar case of antibiotic-resistant bacteria, where genetic mutations give rise to resistance, many adaptations in treatment-resistant cancers are not the result of mutation. The DNA may stay the same, but cancer cells adapt to therapy and outsmart the drugs by switching their gene activity.
Such adaptations do not affect the DNA itself, a hidden layer of regulation controlling the activity of genes — epigenetic and metabolic signals — help cancer cells survive. By targeting this hidden program, it may be possible to overcome deadly cancer resistance.
In new research now published in BMC Systems Biology, the cancer systems biology team of Professor Fabian V. Filipp at the University of California Merced introduces a cellular model of cancer resistance. Under the selective pressure of mitogen activated kinase inhibitors cells are completely rewired yet manage to escape the anti-cancer treatment.
Probing melanoma genomes reveals new targets and challenges
Many patients respond very positively at first. However, unfortunately many ultimately develop resistance and metastases.
To understand how cancer becomes treatment resistant, it is useful to compare pathways in treatment-responsive and treatment-resistant cancers such as melanomas. Melanoma is a cancer that originates in melanocytes, the cells that produce the skin-color pigment melanin and is usually caused by UV damage. Though not the most common form of skin cancer, it is the most aggressive. And if it’s not caught and treated early, it’s also among the deadliest.
UV damage gives rise to point mutations — changes in a single letter of the 3 billion letter human genome. These mutations can interfere with signals that tell cells when to grow and divide and when to stop. Mutations in a protein called BRAF, a major signaling regulator, cause growth signals to be stuck in the “on” position and drive cancer development.
Although scientists have managed to come up with drugs that target and turn off aberrant BRAF signaling, cancer cells are clever and learn to adapt to these BRAF-inhibitors. Today, many patients respond to targeted treatment against BRAF signaling very positively at first. However, unfortunately many ultimately develop resistance and metastases.

Rewiring of gene expression and metabolism to bypass drug blockage
Melanoma is able circumvent BRAF inhibitors not by changing the genes themselves, but by changing gene activity, making them more or less active. Some of the genes with reduced activity were in the same or related signaling pathway as BRAF, the mutated protein that gave rise to the cancer and the main target of chemotherapy. The resistant cells managed to evolve in a way to bypass the signaling blockade or come up with a new way to maintain proliferation. Genes with increased activity were in metabolic pathways that allowed cancer cells to bypass BRAF altogether and continue to grow and divide. Cancer cells had essentially figured out how to survive by rewiring their metabolism in response to chemotherapy.
Systems biology predicts adaptive responses
Daunting as this may sound, it actually offers hope to scientists and clinicians who want to treat chemo-resistant cancers. Non-genomic drug adaptation can be accomplished reproducibly in cultured cells, and combination therapies that block adaptive mechanisms in vitro have shown promise in improving rates and durability of response. Thus, better understanding of mechanisms involved in drug adaptation is likely to improve the effectiveness of melanoma therapy by delaying or controlling acquired resistance.
A better understanding of mechanisms involved in drug adaptation is likely to improve the effectiveness of melanoma therapy by delaying or controlling acquired resistance.
Cancer models in combination with systems biology will pave the way for diagnosis, treatment, and prevention
The systems biology approach reveals that signaling pathways undergo major rewiring at the transcriptional level while acquiring inhibitor resistance. The outcome of such transcriptional plasticity is dysregulation at the level of different upstream master regulators, while maintaining similar effector genes. Combination therapies including targeted approaches and immune checkpoint inhibition are promising and rapidly improving.
For these therapies to show durable, progression-free successes in the clinical setting, adaptation mechanisms of treatment resistances need to be understood. Cellular models in combination with systems biology analyses across epigenomes, transcriptomes, proteomes, and metabolomes provide insight into how non-genomic drug adaptation is accomplished.
Ongoing efforts are focused on utilizing the established preclinical models to overcome drug adaptation as well as precision medicine profiling of cancer patients. Over time, a better understanding of mechanisms involved in drug adaptation is likely to improve the effectiveness of melanoma therapy by delaying or controlling acquired resistance.
Comments