Written by Ahmed B Hamid, Thomas Liehr*
Comment on: Design and validation of a pericentromeric BAC clone set aimed at improving diagnosis and phenotype prediction of supernumerary marker chromosomes; Castronovo et al., Molecular Cytogenetics 2013, 6:45
Castronovo et al. (1) suggest an excellent probe set for the characterization of the centromere-near regions of all human chromosomes. This probe set is an important tool especially for the characterization of small supernumerary marker chromosomes (sSMC), as also outlined in (1). The probe set presented is a very good supplement for aCGH characterized sSMC with cryptic mosaicism (2). Also in case sSMC are present in such low mosaic rates that they may not be characterized by aCGH (3) this probe set might be helpful; also it can be used for larger derivative chromosomes with centromere-near break-events (including balanced rearrangements), as done already with comparable probe sets (4).
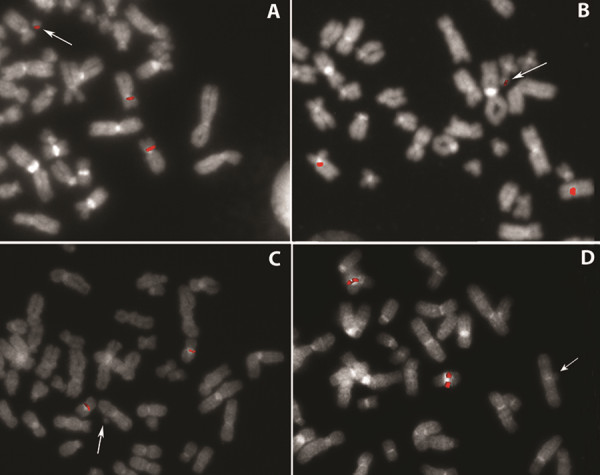 Still, for sSMC we suggest, a clinically relevant pericentric probe set should be generated based on the thoughts outlined in the following. It is known that euchromatic imbalances induced by sSMC may cause clinical problems, but they not always do. Thus, it was hypothesized that centromere-near imbalances only then lead to clinical problems if the concerned region contain dosage sensitive genes (5). Regions being obviously free of such dosage sensitive genes and those containing such genes were narrowed down for most of the pericentric regions already (6-7). This knowledge should be included in an sSMC-specific pericentric probe set.
Still, for sSMC we suggest, a clinically relevant pericentric probe set should be generated based on the thoughts outlined in the following. It is known that euchromatic imbalances induced by sSMC may cause clinical problems, but they not always do. Thus, it was hypothesized that centromere-near imbalances only then lead to clinical problems if the concerned region contain dosage sensitive genes (5). Regions being obviously free of such dosage sensitive genes and those containing such genes were narrowed down for most of the pericentric regions already (6-7). This knowledge should be included in an sSMC-specific pericentric probe set.
We established such a probe set (unpublished data) and showed that it is suited for distinguishing between sSMC leading to clinical problems. For the pericentric region of chromosome 1p for example, it is known that the region free of dosage sensitive genes includes the stretches between 115.8 Mb down to the centromere starting at 121.1 Mb (NCBI 36.3/hg18); for the long arm of chromosome 1 such a region was not defined yet. Also it was shown that sSMC-induced trisomy of the euchromtic region 115.3-121.1 Mb of chromosome 1 lead to clinical problems. Thus, we selected 5 BAC probes spanning the region 114.4-118.5 Mb and another 5 BACs spanning 143.6 to 148.0 Mb starting from cytoband 1q12-q21.1. Similar probe sets are available now for the pericentric regions of all human chromosomes.
A combination of BACs used in the present paper of Castronovo together with our probe set may be the best way to characterize sSMC in a clinically relevant way now and in future.
References:
- Castronovo C, Valtorta E, Crippa M, Tedoldi S, Romitti L, Amione MC, Guerneri S, Rusconi D, Ballarati L, Milani D, Grosso E, Cavalli P, Giardino D, Bonati MT, Larizza L, Finelli P: Design and validation of a pericentromeric BAC clone set aimed at improving diagnosis and phenotype prediction of supernumerary marker chromosomes. Mol Cytogenet 2013, 6:45.
- Hamid AB, Kreskowski K, Weise A, Kosayakova N, Mrasek K, Voigt M, Santos Guilherme R, Wagner R, Hardekopf D, Pekova S, Karamysheva T, Liehr T, Klein E: How tonarrow down chromosomal breakpoints in small and large derivative chromosomes – a new probe set. J Appl Genetics 2012, 53:259–269.
- Liehr T, Klein E, Mrasek K, Kosyakova N, Guilherme RS, Aust N, Venner C, Weise A, Hamid AB: Clinical impact of somatic mosaicism in cases with small supernumerary marker chromosomes. Cytogenet Genome Res 2013, 139:158–163.
- Weise A, Rittinger O, Starke H, Ziegler M, Claussen U, Liehr T: De novo 9-break-event in one chromosome 21 combined with a microdeletion in 21q22.11 in a mentally retarded boy with short stature. Cytogenet Genome Res 2003, 103:14–16.
- Hamid A, Weise A, Voigt M, Bucksch M, Kosyakova N, Liehr T, Klein E: Clinical impact of proximal autosomal imbalances. Balkan J Med Genet 2012, 15:15–22.
- Liehr T: Benign & Pathological Chromosomal Imbalances, Microscopic and Submicroscopic Copy Number Variations (CNVs) in Genetics and Counseling. 1st Edition. 2014. Academic Press.
- Liehr T: Small supernumerary marker chromosomes. https://www.fish.uniklinikum-jena.de/sSMC.html (accessed 11/25/2013).
Acknowledgements: Supported in parts by the DAAD.
*Jena University Hospital, Friedrich Schiller University, Institute of Human Genetics, Germany. Correspondence: thomas.liehr@med-uni.de
Sam Rose
Latest posts by Sam Rose (see all)
- Raising funds for genetic diseases - 23rd September 2016
- The Epigenetics and Chromatin Clinic - 9th November 2015
- Resurrecting one of the oldest genetics journals - 23rd October 2015
Comments