
Single-cell RNA sequencing (scRNA-seq) is a frontier research area that studies genomic events down to the individual cell level, providing unprecedented insights into many biological and medical questions, such as cancer heterogeneity and tumor microevolution. This technology has unique challenges, due to the very tiny amount of starting biological material (RNA) obtained from each cell. As a result, scRNA-seq data are much more noisy compared to the counterpart of conventional RNA sequencing, in which RNA is extracted from a bulk population of cells and is thus much more bountiful. Moreover, as this technique gains popularity the demand for scRNA-seq data analysis has grown exponentially in recently years, posing serious challenges to genomics scientists with limited bioinformatics experience.
We have developed Granatum, a new graphically-interactive web tool that aims to empower scientists with little or no programming experience to analyze scRNA-seq data independently. Granatum is the Latin word for pomegranate, whose little translucent seeds vividly resemble individual cells in scRNA-seq. The website is freely available online here. Without requiring a single line of code, it assists the users through a series of analysis procedures below:

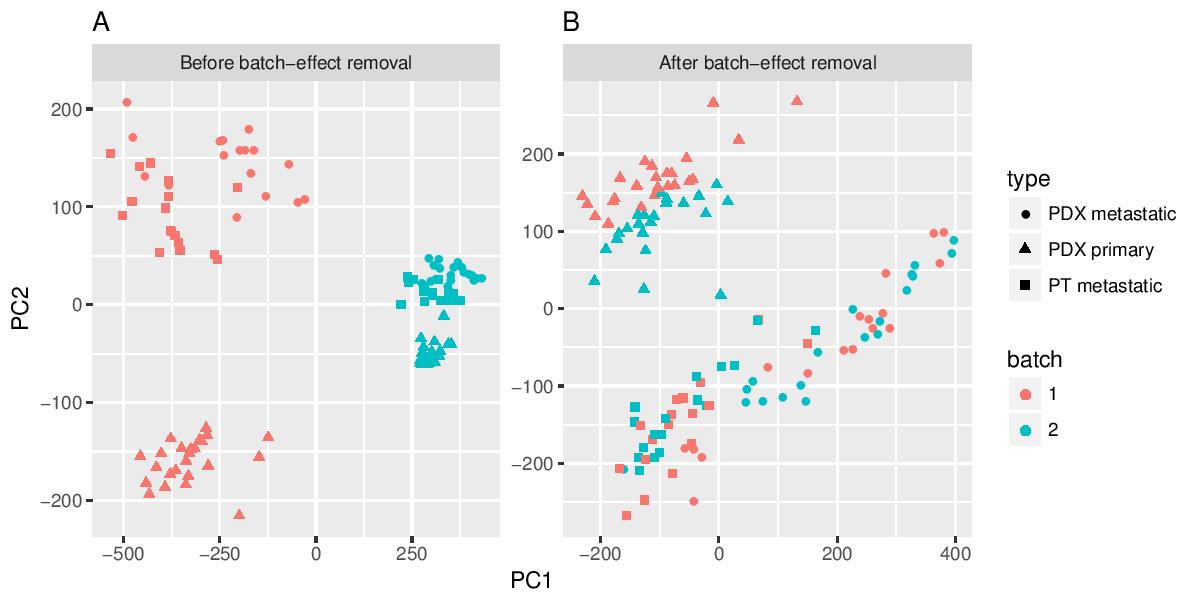
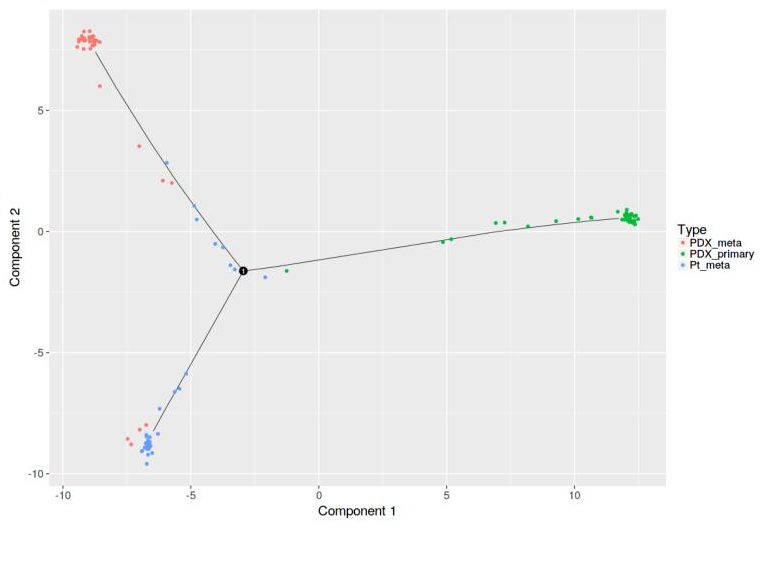
Compared to other tools developed for similar purposes, Granatum stands out with the most comprehensive list of analytical functions (modules), from batch effect removal (A and B) all the way down to reconstructing the developmental trajectories under all single cells (C).
It is also unique for its friendly graphic user interface, providing users with many downloadable and colorful plots and interactive graphs. The development team has added an online user manual and YouTube demonstration video, and can respond to user questions immediately. As a testimony of its utility, this nifty web tool has attracted hundreds of users world-wide at the time of publishing.
As this is a quickly developing field, the Granatum team actively updates the tool for faster speed, broader user base and more comprehensive coverage of analysis methods. Currently plugin standardization is also underway, to make Granatum the tool that truly connects end users and other single-cell bioinformatics developers.
Read the paper: Granatum: a graphical single-cell RNA-Seq analysis pipeline for genomics scientists Learn more about Dr. Garmire’s research group
Comments