Upon this gifted age, in its dark hour,
Rains from the sky a meteoric shower
Of facts . . . they lie unquestioned, uncombined.
Wisdom enough to leech us of our ill
Is daily spun; but there exists no loom
To weave it into fabric.
Edna St Vincent Millay
“Big data”—once the domain of genomics—is now easily the domain of science in general. With new techniques to measure the brain and brain activity (fMRI, EEG, etc) gaining momentum, in the neurosciences tremendous amounts of data are now being generated. The question now is, as Millay points to, how to weave this data into meaning.
During London Technology Week Sean Hill, a co-director of the Human Brain Project (a European brain initiative with €1.2 billion in funding), spoke on this and how this new data-driven collaboration planned to tackle this. “We know a lot about genetics and a lot about high-level cognitive processes,” says Hill. “What we’re missing is the link between the two. How do you go from genes to behavior? That’s the huge challenge.”
A 10-year initiative launched in October of 2013, the Human Brain Project (HBP) comprises a consortium of 256 researchers. The project’s aim is to build, and maintain iteratively, a simulation of the human brain in silico, on a supercomputer, as a resource mapping every neuron and how they connect to one another. At present it is impossible to observe activity within a small group of neurons while simultaneously observing the activity of the entire brain (via imaging technology). A virtual brain would make such observations possible, amongst other things. It would also mean the opportunity to replicate previous in vivo experiments as well as provide a better understanding of the way drugs act on the brain.
Building this virtual brain, however, will be no easy task and running it will mean rethinking our current understanding of computers.
Predictive Build
There are 2,970 possible synaptic pathways in a cortical microcircuit alone. To date, only 22 have been characterized. Mapping these pathways out will be impossible without being able to identify principles in order to predict the rest. But can we accurately predict them? A study by Hill et al in PNAS in 2012 comparing statistical structural connectivity with functional synaptic connectivity provides evidence that for most connection pathways the position for synapses is about the same in vitro and in silico. Because the human brain is the most difficult organism from which to extract data, much of this prediction will be based on the mouse brain. One strand of the project is to generate strategic multi-level data for humans that parallels the data collected for mouse and facilitate the use of mouse data to predict human data.
A New Supercomputer
Building the human brain, of course, is only part of the mammoth scale of innovation this project hopes to accomplish. To then “run” this simulated brain, they will need supercomputing to the exa-scale. Currently, the supercomputers that exist function on the peta-scale. (Indeed, with Europe’s Human Brain Project and Obama’s BRAIN initiative, which estimates yottobytes of data, the neurosciences will easily far exceed genomics as a data-intensive science.)
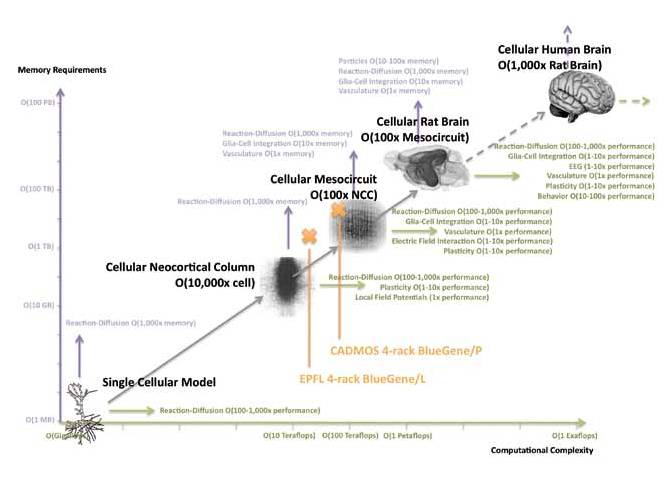
This project builds on the Blue Brain Project, which created a simulation of the neocortical column, incorporating detailed representations of 30,000 neurons. To get some perspective, a similar simulation of a whole brain rat model would include around 200 million neurons, requiring around 10,000 times more memory.
Simulating the human brain would require yet another 1,000-fold increase in memory and computational power. In addition, the creation of virtual instruments (virtual fMRIs, etc) and subcellular modelling would require further computational power (see figure).
To do this, they are working with IBM to build a neuromorphic computing system (see video below)—and aim to have this ready by 2023. This new system will include 4 million neurons and a billion synapses. Traditional computers work through programs (or sets of instructions) and a hierarchy of storage elements telling them what to do in what order and where to find and put things. They are exact and reliable, but in doing this they take up a lot of memory. In contrast to this, a neuromorphic computing system aims to operate like the brain—relying on its ability to learn rather than run through a set of instructions or programs. According to the HBP, “Their architecture will not be generic—it will be based on the actual cognitive architectures we find in the brain—which are finely optimised for specific tasks. Their individual processing elements—‘artificial neurons’—will be far simpler and faster than the processors we find in current computers.”
Open Data
As you might have noticed this new initiative is more of a data integration and infrastructure project than a data generation project. To build this, the project will largely have to take from the data already out there—and there’s lots out there. The way science is published to date, however, will be one of this project’s largest stumbling blocks. To integrate the current data on the brain, more methodological detail on how that data was generated will be needed than what currently constitutes the norm in scientific publishing. In depth knowledge of the experimental protocols, measuring devices, parameters, and so on used to generate the data will be needed to know which data can be reused in building this virtual brain.
This history of poor data management and transparency has real effects. In the clinical setting, increasingly it is becoming apparent that one treatment does not fit all. We will therefore need more personalized treatment to combat some of the most difficult diseases of the mind. However, Pharma are giving up on investing in treatment for brain. Partly for funding reasons, partly because the major findings are not replicable, making the field very risky.
Unlike in genomics where data output tends to be fairly standardised due to the limited number of platforms and thus resulting data types, the neurosciences have operated at a much more specialized, community-driven level, using their own tools and platforms. Open data efforts are, however, starting to emerge through organisations such as the International Neuroinformatics Coordinating Facility (INCF). For example, as our editor Scott Edmunds noted in a blog about neuroscience data notes and our journal GigaScience, one tool promoted by the INCF has been the fMRI data center (fMRIDC)—a large-scale effort to gather, curate, and openly share fMRI data.
A key concern with data sharing across the sciences is the matter of incentive. According to Hill, the neurosciences will have to become an open data endeavour in order to validate future work. Indeed, using predictive modelling to build this virtual brain, the project will need to validate itself through detailed documentation itself of methods and through collaboration and reuse or testing of its findings. It will demand a new more reproducible publishing model—one that includes open data. Hill himself called for a new publishing model, pointing to a future of “living review articles” built using working ontologies (allowing for semantic search of data).
The current publishing model will need to evolve from one based on quantity and proprietorship to one based on quality and reusability. This will demand rethinking current scientific incentives. Hill pointed to a scoring system that could reward organizations that open up their data to the scientific community with more grant money. Data Notes, or short articles specifically focused on the validation of data and its reusability, have also proven a successful open data incentive for our journal GigaScience. This outlet allows for the peer-review of data and thus its validation as well as credit for data through citation.
* * *
The Human Brain Project is one of the few initiatives pointing to a new way of doing science: A science in which data as a default will adhere to the appropriate standards to make it reusable. While there has recently been some criticism of the project and whether we are indeed ready to build a whole-brain simulation, if successful, the project will put in place tools that will help us to know where to look next and which research questions to prioritise. Regardless, it has helped to start a discussion on the very need for more open data and better data infrastructure in the neurosciences. It looks to be an open future for neurosciences.
Just a short clarification over fMRIDC. It did a great job in the past, but due to difficulties in getting funded it has not been available for over a year. I’m sure INCF acknowledges the important role fMRIDC played in the early years of datasharing, but I doubt it is recommending it right not (since fMRIDC has not been accepting submission for quite a while). Luckily the https://openfmri.org is taking over the baton accepting task based fMRI datasets. I recommend can recommend it for all authors wanting to deposit their task based fMRI data (this is what we did as well https://openfmri.org/dataset/ds000114).