
Recognized experts are often vital to bringing the latest findings to a wider audience. In biomedical research, identifying opinion leaders who can raise the visibility of new evidence is increasingly important to how research is communicated.
Traditionally researchers in the field of medical informatics have used methods like surveys, literature searches and obtaing information from other experts to identify possible opinion leaders. Once identified, researchers then use text mining; the method by which a computer can scan plain text to pull out relevant keywords, to assign topics of expertise to the identified names. However these approaches are often time consuming and have the potential to introduce bias when ranking names in terms of importance.
In a new article published in the Journal of Biomedical Semantics, authors Jonnalagadda et al describe a new method for using text mining to both locate and rank potential opinion leaders. Using a collection of articles on obesity, the authors utilized a machine learning system programmed to recognize the presence of keywords such as ‘Dr’, ‘published’, ‘hospital’ etc. 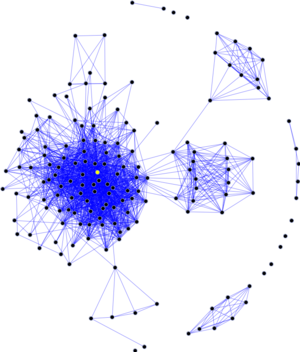
Having compiled a list of subject experts the authors then generated links between persons mentioned in the same news article. This ‘co-mention network’ was then analysed using Social Network Techniques assessing the experts ‘centrality’ or importance in relation to their network.
This interesting use of text mining methods and technology highlights another feature of what is becoming an increasingly promising method of utilizing publically available data for scientific research. With potential applications in genomics and pharmaceuticals, text mining is becoming an essential part of a researchers’ tool box.
Critics of certain publisher’s attitudes to text mining point out that even when permission is granted, text mining is still only possible within that particular website, making text mining’s ability to trawl the entire literature virtually redundant. Funding bodies are also voicing concern, pointing out that as a great deal of research is publically funded, the public has the right to make the most efficient use of any resultant data as possible. In a report launched this week JISC, an established funder of scholarly communications research, comments that text mining opportunities are “hindered by a range of economic-related barriers including legal restrictions, high transaction costs and information deficit which is strongly indicative of market failure.”
There are some publishers, and BioMed Central is one of them, who do act as good text mining citizens, utilizing open access licenses which allow text mining as long as the data is properly attributed. But for the potential of the technology to truly be explored many more scientific publishers will need to follow suit.
The article by Jonnalagadda et al offers a fascinating insight into what may be possible in the field of biomedical research as a result of text mining. More dialogue is now needed between researchers and publishers to develop the kind of licence agreements that will allow this promising technique to be examined. Find out more about BioMed Central’s policy on text mining here. If you wish to join in the discussion about text mining and open access publishing please leave a comment.
Comments