2016: An Eventful Year for GigaScience
Cheering Ourselves up with CUDDEL(s) and Hackathons
We have been working closely with the Metabolomics community for a few years now, participating in hackathon events including BYO Data parties and Hack-the-Spec – ISA as a FAIR research object – thanks to our collaboration with the ISA Team at the Oxford e-Research Centre and funding from our BBSRC UK-China partnering award. Following the success of these events, we were awarded a second UK-China collaboration grant. On top of working again with the ISA-team, this time we have partnered with the EBI, the Universities of Birmingham, Manchester and Oxford, The Sainsbury Laboratory and the Earlham Institute (formerly TGAC). The CUDDEL project has a focus on the ISA framework for describing data generated from research experiments.
#metabolomics #opendata #reproducibility some of the topics in the https://t.co/ua1rsMsDMB hackathon, here some of the participants pic.twitter.com/VAYd4ixJCn
— AlejandraGonzalezBeltran @alegonbel@mastodonapp.uk (@alegonbel) November 25, 2016
The GigaScience technical team attended the latest CUDDEL workshop which was hosted by Chris Steinbeck and his team at the EBI during the last week of November. With help from our CUDDEL collaborators, we have begun using ISA’s application programming interface in GigaDB so that metadata descriptions of datasets are downloadable in ISA format. We are also currently working with the authors of a human metabolomics study on the effects of food intake and exercise, facilitating its reproduction using ISA and computational workflows. We hope to make both these resources available in early 2017 and add to the papers we are currently gathering in our Integrative Metabolomics Series.
Community Curation and the Annotometer
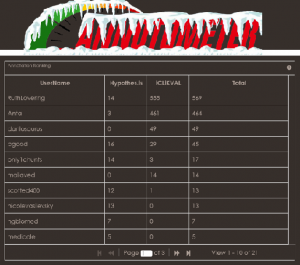
GigaScience is keen to maximize the use and utility of our published data, narrative and research objects, in which curation plays a key role. However, curating the global scientific record it is an impossible task for one or small group of professional curators to achieve, so we thought the power lies in the hands of people who use the data. At this year’s Biocuration 2016 meeting we decided to roll up our sleeves and try to encourage community annotation of our and others data resources through a “GigaCuration Challenge”. The competition was in collaboration with two open annotation tools – Hypothes.is and iCLiKVAL; we also developed a new tool, the Annotometer – a live leader board, specifically for this challenge. An iPad was the grand prize awarded to Ruth Lovering (UCL), who made the most annotations. Also, as GigaScience is part of the Annotating All Knowledge – Hypothes.is working group, our Editor, Nicole, presented the results and lessons learned as a use case at the FORCE 2016 meeting.
We have made the code for the Annotometer open source and we would like to encourage everyone to run their own annotation competitions and to use our leader board – help us build upon the code to improve the tool.
Reproducible Parasites with Protocols.io
 We thought we were at the very frontline of reproducible research publication until we collaborated with Protocols.io – a novel platform for sharing methods than can be cited and updated over time that serves to improve reproducibility even further. Our new collaboration with Protocols.io was launched with the publication of two research papers on scabies and tapeworm (Schistocephalus solidus) that demonstrates a new way to share scientific methods that allows scientists to better repeat and build upon complicated studies on difficult-to-study parasites. It’s great to see these authors (and others in our pipeline) take advantage of Protocol.io’s open access repository of scientific methods and collaborative protocol-centered platform. Taking things even further, working with the protocols.io team we’ve become, the first journal to provide deep integration into the submission, review and publication process. We now also have a groups page on the portal where our methods can be browsed. So, watch this space as we publish more papers using Protocols.io.
We thought we were at the very frontline of reproducible research publication until we collaborated with Protocols.io – a novel platform for sharing methods than can be cited and updated over time that serves to improve reproducibility even further. Our new collaboration with Protocols.io was launched with the publication of two research papers on scabies and tapeworm (Schistocephalus solidus) that demonstrates a new way to share scientific methods that allows scientists to better repeat and build upon complicated studies on difficult-to-study parasites. It’s great to see these authors (and others in our pipeline) take advantage of Protocol.io’s open access repository of scientific methods and collaborative protocol-centered platform. Taking things even further, working with the protocols.io team we’ve become, the first journal to provide deep integration into the submission, review and publication process. We now also have a groups page on the portal where our methods can be browsed. So, watch this space as we publish more papers using Protocols.io.
Our Coming of Age and DORA (not the explorer)
The annual Intelligent Systems of Molecular Biology (ISMB) meetings hold a special place in our heart – we’ve celebrated our birthdays at these events in what seems to becoming a tradition – a great opportunity to catch up with numerous new and old friends. And this year was no different, we celebrated our coming of age (4th birthday) at DisneyWorld in Orlando. Unfortunately, the aging process isn’t all pretty, and another sign of getting older was the acquirement this summer of our getting indexed in a notorious database that has a glaring negative impact on research and scientific communication. Not wanting to give it any more attention than it deserves, our Editor-in-Chief’s answered people’s questions with a last word on the problems with the journal Impact Factor here. And to help us take a stand against it – we recommend, like us, that you sign DORA (Declaration of Research Assessment) and follow their recommendations for reducing its use.
Parties and Fighting the Zombie Paper Apocalypse
Last month, we celebrated Neuro-November – our Editor, Nicole, had a lot of fun in San Diego, where she attended the SfN 2016 (Society for Neuroscience) meeting and we collaborated with some of our FORCE11 open science and reproducibility partners, NIF (Neuroscience Information Framework), SciCrunch, the Resource Identification Initiative, and Hypothes.is, in an annotation competition, “Battle Against the Zombie Paper Apocalypse”. The goal of the competition was to help neuroscientists bring their dead, static papers back to life by making them more reproducible by annotating (via the Hypothes.is tool) a research resource, such as animal models, software, cell lines or antibodies, with an RRID (research resource identification). Competitiveness against different Universities was encouraged through further use of our Annotometer leader board (see above), and we had over 100 papers annotated and made a step closer to being more reproducible. Once again our Giga-swag (t-shirts, stickers and Panda USB sticks) proved extremely popular, and were given as prizes to anyone who went one step further – by either signing up for a Hypothes.is account or registering a new resource in SciCrunch.
The zombie paper apocalypse battle is taking off and leaderboard is live! #sfn16 @neuinfo @hypothes_is @SciCrunch pic.twitter.com/WEHXKEizMZ
— GigaScience (@GigaScience) November 13, 2016
The @INCForg @neuinfo @GigaScience @SciCrunch party was a success! @RCCraddock and @memartone pushing for #openscience #sfn16 pic.twitter.com/0tjzozDvNQ
— GigaScience (@GigaScience) November 15, 2016
However, it wasn’t all about fighting the Zombie papers, GigaScience also co-sponsored a party and SfN Banter on the theme of “Reproducible Research through Open Science” in conjunction with INCF, NIF, SciCrunch, Mendeley and CaymanChem. The party featured two lightning talks from GigaScience Ed Board Members, Cameron Craddock and Maryann Martone – who pushed for more open science and reproducibility with RRIDs. It was a fun night meeting old and new friends over Zombie Drinks, Negronis and more – with a final turnout of 1000 people! We’ll definitely need a bigger venue next year.
Open BIG Data – the Gift that Keeps on Giving
Welcome To the club Mr. Olive, genome sequenced! @toni_gabaldon @gabaldonlab @CRGenomica https://t.co/6bgZ4BLuvM pic.twitter.com/XBnz8iaUcH
— Sketching Science (@sketchscience) June 29, 2016
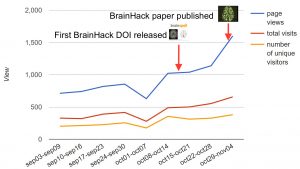
This year we are proud to have published and released such a variety of different big data types for people to re-use and build upon. Some of this year’s highlights include publishing the genome of an ancient 1,200-year-old olive tree – finally joining the genome club with its related data in the public domain. We also brought you Apple genome 2.0 – in a new long-read format, as well as 425,000 camera trap images of California desert animals. We also published the genome of the world’s largest bony fish, the ocean sunfish (Mola mola)–a project initiated by Nobel Laureate, Sydney Brenner, as well as data from the Personal Genomes project in a new phase, led by the man with the largest lab at Harvard, George Church. also bingeing on sweet brains (research outputs) with the publication of skull-stripped anatomical MRI data and the Brainhack America’s project reports and their associated code and metadata in GigaDB, with each project having their own citable DOI. Like our early release of the datasets of the Avian Phylogenomic Project, the staggered early release of these projects in GigaDB and social media increased traffic to our repository over 50%.
Last but not least, our last paper to be published by co-publisher, BioMed Central (BMC), was the genome of the “living fossil”, Gingko Biloba – ending our time with BMC on a high, this paper was featured in The Guardian and BBC News.
Goodbye BMC – Hello OUP
Some of our readers may already know, but for those of you who don’t, our final big news to end the year is that GigaScience will be moving publisher. From 1 January 2017, we will be published by Oxford University Press (OUP). After almost 5 years at BMC, we would like to take this opportunity to thank them and in particular, all the staff whom we have worked with closely. You can read more about our move to OUP in the press release. Our move will only mean more exciting and eventful things in 2017 and beyond, so watch this space as we continue to push the boundaries of innovative open science publishing. And with that, we would like you all to have yourselves happy and well-earned holidays, and we will too.