Parkinson’s disease (PD) is a common neurodegenerative disorder with a prevalence of 0.5-1% among people over 65 years of age. More than 10 million people worldwide currently live with PD.
Unfortunately, PD is hard to diagnose accurately in the early stages of disease for the following reasons:
1) The characteristic motor symptoms – rigidity, slowness of movement and tremors – do not develop until 50-80% of the dopaminergic neurons in the mid-brain – the source of dopamine in the brain –are lost. Dopamine deficiency results in some of the motor features in PD. Before the onset of these characteristic symptoms – during what is called the prodromal phase that can last for a long time – many nonspecific features of PD dominate.
With 831 subjects in total, our study is the largest yet into the possible role of DNA methylation in PD
2) Other disorders such as post-traumatic, drug- or toxin-induced and arteriosclerotic disorders share the symptoms of PD.
3) Motor and non-motor symptom progression is very variable and unpredictable.
Because of these difficulties, there is an urgent need to develop early diagnostic or prognostic biomarkers for PD that allow for PD to be identified before the onset of symptoms.
Recently, there has been a growing interest in the role of DNA methylation in PD. DNA methylation is an epigenetic mechanism that regulates gene function through the addition of a methyl group to cytosine – one of the building blocks of DNA – without altering the DNA itself. Abnormalities in DNA methylation have been associated with a number of health conditions.
With 831 subjects in total, our study is the largest yet into the possible role of DNA methylation in PD. It is more than 4 times larger than any similar studies that have been published so far and unique in that we used a population-based design to recruit PD patients and controls. –We combined the “Parkinson’s disease, Environment and Genes (PEG)” study with the highly robust Illumina array platform – a methylation screening tool – to generate genome-wide DNA methylation data.
Our findings serve as a proof of concept that tissues such as blood and saliva – which are easy to sample – can be useful in finding PD biomarkers
Using both epigenome-wide association and network analyses – a novel approach in epigenome-wide studies in PD – to investigate the association between PD and DNA methylation changes by site or as networks (clusters), we show that PD status is indeed associated with DNA methylation changes in two tissues: blood and saliva. Many of the differences in methylation levels were due to different amounts of various types of white blood cells found in the blood of PD patients compared with controls which implicates the immune system in the development of PD.
Our findings serve as a proof of concept that tissues such as blood and saliva – which are easy to sample – can be useful in finding PD biomarkers.
We also analyzed certain biologic processes within a cell or tissue known as pathways –– to provide the first evidence that five of these pathways are different in PD patients compared with controls: cell energy generation (mitochondrial function), axonal transport , systemic immune response, cell signaling through Wnt involved in development and self-renewal of tissues, and brain iron handling.
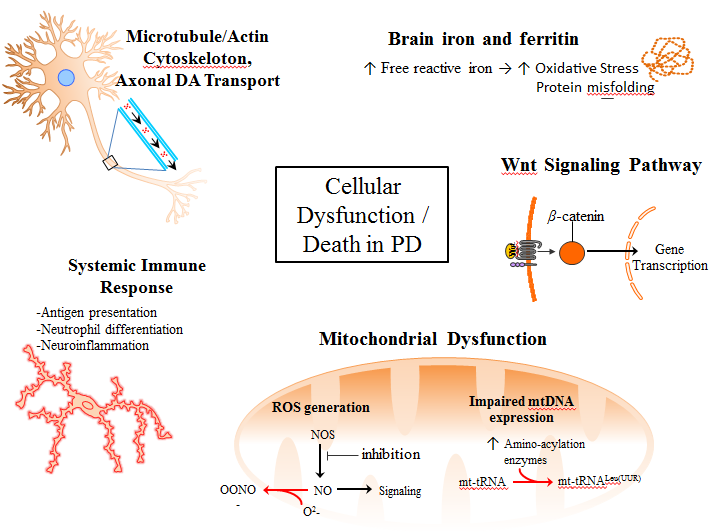
Our results confirm that previously suggested biological pathways are indeed involved in PD. We also provide additional evidence for the involvement of previously underappreciated pathways.
Our study is helpful for the development of biomarkers for diagnosis, prognosis, and targeted therapy of PD. Additionally, our findings highlight the role of the immune system in PD. Nevertheless, future research will need to address the causal structure of these relationships.
Comments