
Wood decomposition under hypoxia: Towards sustainable bioethanol production – Hans Mattila
Basidiomycota fungi that grow on tree trunks and pieces of dead wood are a familiar sight in our forests. However, the eye-catching fruiting body of a wood decomposer is only part of the organism. Wood decomposition itself is actually orchestrated by its hypha that elongates beneath the bark and even penetrates the dead wood.

As a mycologist, I wondered how wood decomposition can take place inside waterlogged pieces of wood where the hyphae must encounter temporal or regional oxygen depletion. This led us thinking that these organisms could be capable of fermenting ethanol while decomposing untreated wood-based materials under hypoxia.
Our research presents new potential regulative factors within the well-studied ethanol fermentation and primary metabolism pathways.
I started my Ph.D. project to test this hypothesis by screening Basidiomycota Polyporales fungi for their capability for simultaneous saccharification and fermentation (SSF). A white-rot isolate Phlebia radiata 79, outperformed other candidates by bioconverting various unpretreated lignocellulose materials under oxygen-limited conditions. P. radiata was capable of decomposing softwood, hardwood, recycled wood waste as well as barley and wheat straw (Mattila et al. 2017, 2018).
We extended our research to better comprehend how SSF related pathways are regulated in P. radiata. Thus we extracted and sequenced RNA from various culture conditions and combined the information with promoter regions analysis.

First, we noticed that P. radiata reacts differently to hypoxia compared to well-studied Ascomycota species. In fact, the hypoxic response of P. radiata resembles the one of a Basidiomycota species Cryptococcus neoformans. Our research presents new potential regulative factors within the well-studied ethanol fermentation and primary metabolism pathways.
Second, we learned that hypoxia changes the profile of secreted wood decomposition (CAZy) enzymes. Based on our results, hypoxia seems to be an equally important regulator of CAZy genes as is the composition of the growth substrate (Kuuskeri et al. 2016).
Altogether we believe our research moves us one step closer towards sustainable 2nd generation bioethanol production.
In conclusion, our research tries to shed light on the regulative aspects behind SSF. This has aided our applicative project which aims to a novel type of bioethanol production method. We try to comprehend the gene regulation of a wood decomposing organism that has the capability to use a versatile selection of solid substrates. Altogether we believe our research moves us one step closer towards sustainable 2nd generation bioethanol production.

Hans is Doctoral Student in the Department of Microbiology at the University of Helsinki, Finland. He started his Ph.D. project to test the hypothesis that Basidiomycota fungi could be capable of fermenting ethanol while decomposing untreated wood-based materials under hypoxia. The research started by screening Basidiomycota Polyporales fungi for their capability for simultaneous saccharification and fermentation (SSF), and then it was extended to better comprehend how SSF related pathways are regulated in P. radiata.
Leveraging metatranscriptomics and synthetic biology to enhance our understanding of the biomass-degrading machinery of anaerobic fungi – Itai Brand-Thomas
I am intrigued by how microbes interact with their host environments and the effects of these interactions on human and animal health. I am excited about the opportunity to conduct interdisciplinary research that utilizes both cultivation independent omics techniques and wet-lab microbiology to learn about a phylogenetic group that is poorly understood, despite its importance to host function and health.
Prior to the mid-1970s it was assumed that fungi required oxygen for survival and growth…
Prior to the mid-1970s it was assumed that fungi required oxygen for survival and growth, and it was only during this time that anaerobic fungi were described as such. Since then it has been established that anaerobic fungi from the rumen represent some of the most prolific known biomass-degraders and recent genome studies have shown that anaerobic rumen fungi have genomes rich in carbohydrate active enzyme (CAZymes). The proper function of the rumen relies on the interaction of archaea, bacteria, fungi, protozoa and viruses.
Whereas the prokaryotic rumen population has been well-researched using cultivation dependent and cultivation independent techniques, the number of studies concerning the fungal population of the rumen remains low. The lack in knowledge of the biology of this important functional group leads to an incomplete picture of the rumen ecosystem, and results in inaccuracies in models that predict how rumen function and health is affected by changes in the ruminants’ diet.
By studying rumen fungi and their enzymes, it will be possible to enhance our understanding of the synergistic nature of the rumen microbial community and to improve current models. This will facilitate the development of advanced strategies that improve rumen function and the health of the host animal while simultaneously reducing the environmental impact of enteric fermentation, namely reducing the production and release of the potent greenhouse gas methane.
Furthermore, I anticipate that previously uncharacterized CAZymes from anaerobic fungi might complement their bacterial counterparts, representing the missing link to develop more efficient enzyme cocktails for industrial biomass degradation and the production of novel bioproducts.
…uncharacterized CAZymes from anaerobic fungi might complement their bacterial counterparts, representing the missing link to develop more efficient enzyme cocktails for industrial biomass degradation and the production of novel bioproducts
For my own research, I have been utilizing metatranscriptomics to study the anaerobic fungi that colonize and degrade recalcitrant plant material during rumen-incubation. I have been able to assemble thousands of CAZyme transcripts from the population of anaerobic rumen population, and subsequent sequence analysis has revealed that these fungal CAZymes have a rather low overall sequence similarity to currently known, primarily bacterial CAZymes. The CAZymes identified in the course of this work is expected to significantly expand the diversity within the CAZyme universe.
I am currently in the process of biochemically characterizing and obtaining crystal structures of some of the newly identified CAZymes to verify their predicted activity and to obtain a better understanding of the molecular mechanisms by which these fungal CAZymes facilitate the degradation of plant biomass under anaerobic conditions.
…find a topic that you are passionate about and use that excitement to fuel your research.
The advice I would give to young scientists starting out in research is: find a topic that you are passionate about and use that excitement to fuel your research. Don’t be intimidated by large amounts of data or a technique you are unfamiliar with; give it a shot, ask questions along the way, and learn from your mistakes. Make sure to find time for activities outside of research. This will allow you to approach your studies with a different perspective and allow you to take a mental break and re-energize.
(Acknowledgements: I was able to attend to the Fungal Genetics Conference thanks to funding from the Alumni Foundation of Amherst College).
 Itai is a first-year graduate student in the Systems Microbiology & Natural Products Laboratory at the University of California in Davis (USA). Her thesis project centers on anaerobic fungi that inhabit the rumen of the cow and the enzymes secreted by this significantly understudied group of microorganisms.
Itai is a first-year graduate student in the Systems Microbiology & Natural Products Laboratory at the University of California in Davis (USA). Her thesis project centers on anaerobic fungi that inhabit the rumen of the cow and the enzymes secreted by this significantly understudied group of microorganisms.
Innovations for the fermentation industry using filamentous fungi – Ken Miyazawa
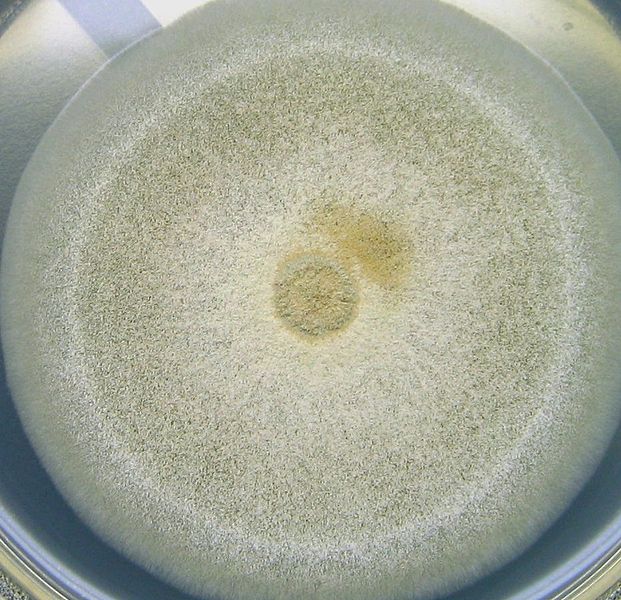

The cell wall of Aspergillus species is mainly composed of polysaccharides including α-1,3-glucan. We previously reported that α-1,3-glucan has a role for hyphal aggregation in Aspergillus nidulans which possesses two α-1,3-glucan synthase genes (agsA and agsB).
Here, we constructed the strains overexpressing agsA (agsAOE) or agsB (agsBOE). The degree of hyphal aggregation was different between the overexpressing strains; tightly aggregated in the agsBOE, whereas loosely aggregated in the agsAOE.
Size-exclusion-chromatography analysis of α-1,3-glucan revealed that the molecular mass of α-1,3-glucan from the agsAOE was four times larger than that from the agsBOE.
In addition, fluorophore-labeling of hyphal polysaccharides suggested that spatial localization of α-1,3-glucan in the cell wall was different between the overexpressing strains; outer layer in the agsBOE, whereas in the inner layer in the agsAOE. Taken together, hyphal pellet formation depends on the molecular mass and localization of α-1,3-glucan as well as the amount of α-1,3-glucan in the cell wall of A. nidulans.
Our research would provide a potential innovation for the fermentation industry that uses filamentous fungi.
Since we previously reported that the α-1,3-glucan-deficient mutant of the industrial fungus Aspergillus oryzae increased twice in the secreted protein production than the wild-type strain, understanding the mechanism of hyphal aggregation could contribute to improvement of fermentation productivity. Our research would provide a potential innovation for the fermentation industry that uses filamentous fungi.

Ken is a PhD-student in the laboratory of Applied Microbiology (Prof. Keietsu Abe) at Tohoku University in Japan. His research project is to develop novel technology that improves fermentation productivity in fungal industry through analyzing function of cell wall polysaccharides.
Development of fungal-selective molecules and strategies to target Hsp90 – Emmanuelle LeBlanc
Fungi infect over a billion people worldwide, posing a great burden on human health. Invasive fungal infections kill an estimated 1.5 million people annually. These numbers continue to rise with the increase in immunocompromised populations and high-risk individuals. The three most prevalent agents of invasive fungal disease are Candida albicans, Cryptococcus neoformans and Aspergillus fumigatus. New therapeutic strategies are imperative as the three major classes of antifungals are hampered by host toxicity, the emergence of drug resistance, and a narrow activity spectrum.
Selective targeting of stress responses in fungal pathogens provides a promising therapeutic strategy to mitigate drug resistance and combat invasive mycoses. The molecular chaperone Hsp90 has been extensively validated as an essential regulator of virulence traits and antifungal resistance in Candida species. However, toxicity of the current Hsp90 inhibitors, that inhibit the host chaperone, impedes their use as antifungal treatments.
My project aims to assess the therapeutic potential of targeting Hsp90 in fungal pathogens. I work closely with a fantastic international team including experts from the Structural Genomics Consortium, and the Center for Molecular Discovery at Boston University. Together, we are optimizing fungal-selectivity in analogs of a natural-product Hsp90 inhibitor by structure activity relationship. This interdisciplinary collaboration has allowed us to combine structural, chemical, biochemical, and genetic approaches to design synthetic compounds with >25-fold fungal-selectivity for C. albicans and C. neoformans.
My goal is to uncover new biology and provide important insights supporting the feasibility of targeting Hsp90 and its regulatory circuit to treat deadly fungal infections.
In parallel, I aim to identify fungal-specific Hsp90 circuitry that governs stress responses and virulence in C. neoformans. My goal is to uncover new biology and provide important insights supporting the feasibility of targeting Hsp90 and its regulatory circuit to treat deadly fungal infections.
 Emmanuelle is a Master’s student in Dr. Leah Cowen’s lab, in the Department of Molecular Genetics, at the University of Toronto. She is also a triathlete and an aspiring clinician scientist.
Emmanuelle is a Master’s student in Dr. Leah Cowen’s lab, in the Department of Molecular Genetics, at the University of Toronto. She is also a triathlete and an aspiring clinician scientist.
When two fungi quarrel the human profits – Norman Paege

I am in the sixth year of my doctoral degree studies at the TU Berlin with two short interruptions due to paternity leave.
My research focus is on the mode of action of the antifungal protein AFP secreted from the filamentous fungus Aspergillus giganteus. It is of great importance to develop new antifungal strategies, particularly due to a world-wide increase in problems with fungal resistance and infections, which has resulted in damage to agriculture and crop loss, and serious impact on human health.
In our research project, we have first tried to find the elements which determine AFP sensitivity. Our focus is on the fungal plasma membrane and the differences between fungal species. We have found that small changes are important and have a large impact on AFP susceptibility.
Secondly, we are working on understanding the role of AFP plays for its host. We have found more than 80 orthologues of this peptide spread throughout the fungal kingdom. Luckily, one, the AnAFP, is produced by A. niger. With the use of our transcriptome database including 155 growth conditions, we concluded that AnAFP is involved in several metabolic processes active during carbon starvation or autophagy.
…it is crucial to define your own topic right at the beginning of your doctoral research, and try to be consistent in following it through.
In my research experience I have learned – and this is the advice I’d like to share with young scientists starting out in research – that it is crucial to define your own topic right at the beginning of your doctoral research, and try to be consistent in following it through.
In addition, another piece of advice I found important: If you want to use a new method, try to start a collaboration with a lab where it is established. This will ensure support and better progress.

Norman is Doctoral Student in the Department for Applied and Molecular Microbiology (AMM) at TU Berlin, Germany. His research focuses on studying the mode of action of the antifungal protein AFP secreted from the filamentous fungus Aspergillus giganteus, and his research interests aim at developing new antifungal strategies against increasingly more resistant fungal infections.
Roberto Garbero
Latest posts by Roberto Garbero (see all)
- Local knowledge systems: how can they help to guide ecological transition and a freer world? - 30th November 2022
- Scientists of the future at the 30th Fungal Genetics Conference - 9th May 2019
- Philosophy of life sciences is ‘constructive subversiveness’ - 16th November 2017
Comments