Non-alcoholic fatty liver disease: What we know and what we don’t
Nonalcoholic fatty liver disease (NAFLD), covers a wide spectrum of liver pathology that are not caused by alcohol consumption, spanning from fat accumulation in the liver (steatosis), nonalcoholic steatohepatitis (NASH), cirrhosis, and hepatocellular carcinoma. NAFLD is the most common liver disease, affecting 25% of the world population, and the prevalence continues to rise along with other closely associated conditions such as diabetes and obesity. Currently, there are no existing therapies directly targeting NAFLD.
The etiology of NAFLD is highly complex and poorly understood. Previous research has found lipid metabolism, inflammation, and a few other processes and genes to be important, but how these genes and processes interact and whether other pathways are involved are unclear.
One particular interesting observation about NAFLD is the clear differences between males and females. At younger ages, males are more likely to have NAFLD than females. Males also demonstrate more severe metabolic dysfunction, such as higher glucose levels, higher blood pressure, greater visceral adiposity, and greater liver injury. However, at older ages, especially after menopause, the prevalence in females increases. The exact mechanisms for these sex differences are unclear.
Using Big Data to answer Big Questions
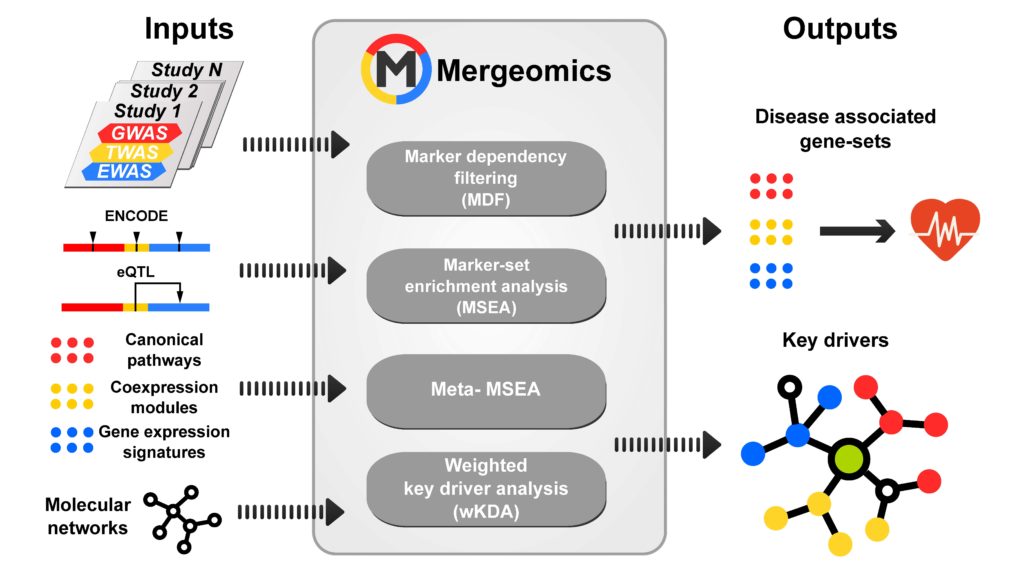
We set out to analyze the ample genomic data gathered around NAFLD using a method my lab has developed, called Mergeomics (read more about Mergeomics in the BMC Genomics paper here). Mergeomics is designed to take diverse types of genomics data (genetic, transcriptomics, proteomics, metabolomics etc) that are associated with a disease or phenotype and integrate them together to map out disease-association signals segregating on biological pathways and gene-gene networks. Mergeomics can also predict important regulatory genes, called “key drivers”, that likely control other disease genes and are therefore more important for the disease.
Using Mergeomics, we analyzed a large mouse dataset generated by our collaborator, Dr. Jake Lusis and his team. This dataset, called Hybrid Mouse Diversity Panel (HMDP), contains genetic information for millions of genetic variants across ~100 mouse strains, gene expression data of tens of thousands of genes from liver and fat (adipose) tissues from hundreds of mice of each sex, and careful measurements of NAFLD characteristics. In a recent study by our two groups, we examined the male mice data and found numerous genes and biological pathways in both liver and adipose tissues to be important for NAFLD, with the majority of them not being discovered before. Dr. Lusis’ group tested several newly predicted genes from our analysis and confirmed their critical role in NAFLD development, thereby highlighting the power of our big data integration approach to uncover novel biology.
Steps towards understanding sex differences in NAFLD
In a recent study published in Biology of Sex Differences, we applied Mergeomics to the female Hybrid Mouse Diversity Panel data and again identified a plethora of NAFLD-associated genes and pathways interacting in both the liver and adipose tissue, many of which are novel. When comparing the results between males and females, we found both shared and sex-specific biological processes for NAFLD. Shared functions included: adaptive immunity, branched chain amino acid metabolism, mitochondria function, and cell cycle. Processes such as vitamins and cofactors metabolism and ion channel transport were specific for females whereas phospholipid, lysophospholipid, and phosphatidylinositol metabolism and insulin signaling were specific for males. Additionally, numerous lipid and insulin-related pathways and inflammatory processes in the adipose and liver tissue appeared to show much stronger association with NAFLD in males than in females. This finding aligns with the more severe metabolic dysfunction (due to NAFLD) that is observed in males.
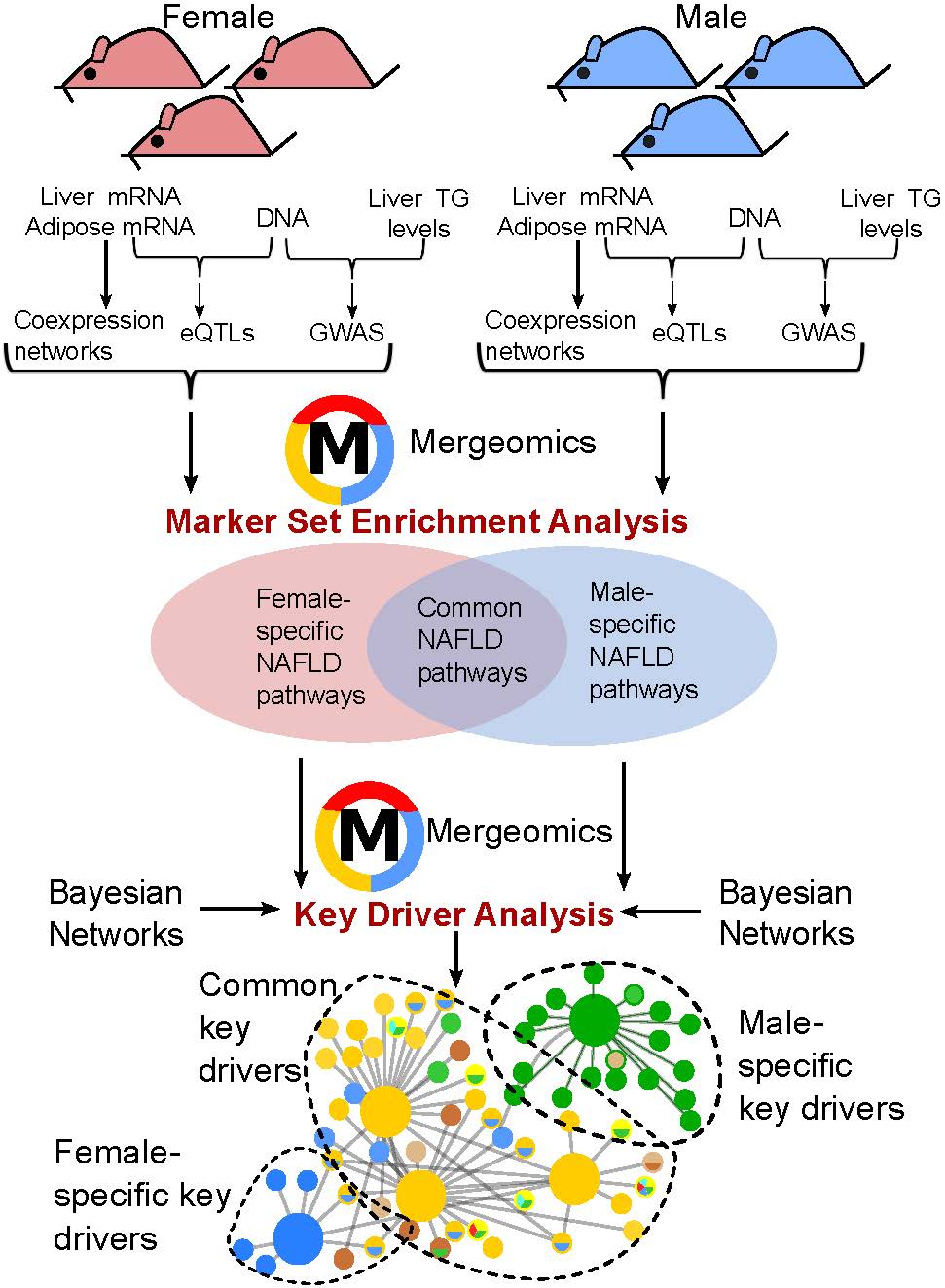
In addition to biological processes, we also predicted key driver genes orchestrating the NAFLD pathways in a sex- and tissue-specific manner. Our research suggests that sex hormones do play a partial role in initiating dramatic sex differences in the pathology of NAFLD.
In summary, this study represents an example of using big data science to tackle complex, yet fundamental, biology questions. There is still a lot more research to be done—such as examining the role of sex chromosomes and experimentally testing the female-specific regulatory genes uncovered by our study to confirm their roles in NAFLD. Nonetheless, our findings shed light on the biological mechanisms driving sex differences in NAFLD and can guide future translational studies to develop novel therapies against NAFLD with greater precision and efficacy for both males and females.
Comments