
Genetics of population isolates
“Population isolates” refer to ethnic groups, which are separated from their surrounding populations by barrier(s) and have minimal gene flow from them. These barriers can be geographical, cultural or linguistic. Moreover, isolated populations also act as a catalyst for studies aimed at understanding the basis of common diseases. Therefore, studying population isolates have always been of great interest for the anthropologists.
Two new papers published in Genome Biology focus on two such isolates and explore their demographic history and genomic selection and while I pen down my thoughts about them, I recall two individuals from these populations who shared their thoughts during a journey. Little did I know then, that the information shared by them would be a connecting thread to write on these population isolates.
Parsi: like sugar in milk

Parsi is one of the smallest ethnoreligious communities of the world, with around 57,264 members as per the 2011 Indian census (down from 69,601 in 2001 and 114,000 in 1940). Their birth:death ratio steadily declining over the past century, thus making them more vulnerable.
To control this decline, in 2013 the Indian Government created the “Jiyo Parsi” (literal meaning to live Parsi) campaign.
This population traces its origin to Persia (present day Iran), from where they fled during 8-10th century following Islamic conquest. A group of Zoroastrian first landed in Sanjan (Gujarat, India), and were granted asylum by the local King named Jadi Rana with a promise to mix culturally with native Hindus “like sugar in milk”.
The Parsi make up less than 0.005% of India’s population, but as described by the famous Indian writer Amitav Ghosh, “they have essentially created modern India”.
Genetic history of Parsi populations
Previous studies on Parsi populations have relied mainly on small sample sizes and low resolution genetic markers, limiting the power of the analysis. Moreover, the major Parsi group (living in India) has been often underrepresented, and likewise, no high-resolution autosomal evidence has been considered in this debate.
Both of the Indian and Pakistani Parsi populations are genetically closer to each other and Iranians than they are to the neighboring Indian and Pakistani populations.
Using a combination of high resolution uniparental and biparental genetic markers on modern Parsi samples as well as mtDNA markers on ancient Parsi samples, a recent study specifically looked at whether modern Parsi people living in India and Pakistan have a common genetic basis, shared among them and also the Iranian population, or if their genomes had been affected by admixture with their Indian and Pakistani neighbors? Also, to know that at what extent current Parsi populations of India assimilated local females in the past? First successful ancient DNA data from remains excavated in Sanjan (Gujarat, India) were analyzed.
Admixture with the local South Asian populations was minimal in comparison to other historical migrants to India (e.g. Jews and Siddi analyzed in our previous work). Both of the Indian and Pakistani Parsi populations are genetically closer to each other and Iranians than they are to the neighboring Indian and Pakistani populations. Neolithic Iranian samples shared higher number of alleles with Parsis than modern day Iranians.
Of course, the modern Iranian population exhibited marked difference from Parsis, in that they carry an additional European ancestry, and significantly lower South Asian specific ancestries. This genomic difference was likely due to Islamic conquest in Persia.
Indian and Pakistani Parsi populations showed high level of homozygosity in their genome compared to their putative parental and present neighbors. This is likely due to small population size and high level of inbreeding.
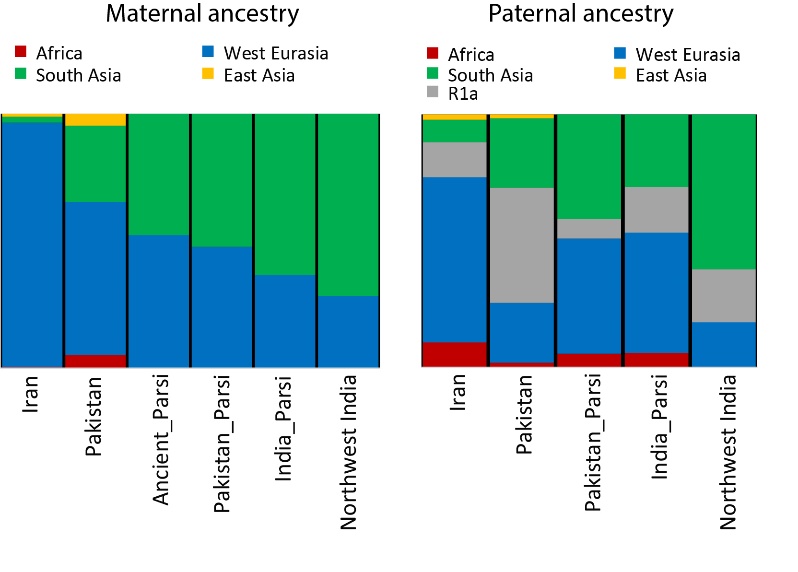
Maternal ancestry was examined via analysis of mtDNA haplogroup composition in the ancient remains excavated from Sanjan. With this information, the authors observed that there seems to have been some assimilation of local South Asian women during initial settlement, indicating a single wave of population admixture that corresponds with the historical date of arrival of Paris to South Asia. On the other hand, paternal heritage aligns between the Iranian and Pakistani populations.
Sherpa and Tibetan: the fine scaled differentiation
People living at high altitude, such as the Sherpa in Nepal, have been exposed to low pressure and decreased oxygen levels for several thousand years, facilitating natural selection to live efficiently under high altitude hypoxia.
Scientists have suggested that high altitude adaptation across the world is not necessarily shared via identity-by-descent. For example, Andean highlanders’ high-altitude adaptation is enabled by increased oxygen-carrying blood hemoglobin, while less blood hemoglobin is a hallmark of Tibetan and Sherpa high-altitude adaptation. The decreased blood hemoglobin helps these populations avoid blood clots and strokes caused by thickening of the blood.
A series of studies on Sherpa and Tibetans have unraveled the mystery of high altitude adaptation layer by layer.
Genetic histories of Sherpas and Tibetans
The genetic origin and interrelation of Sherpa and Tibetan has been a topic of hot debate recently. Studies using sex-specific markers (mtDNA and Y chromosome) have suggested that the Sherpa population is a recent offshoot from Tibetans, while studies based on autosomal data and high altitude adaptation find that the Tibetan population originated following admixture between ancestral Sherpa and Han Chinese.

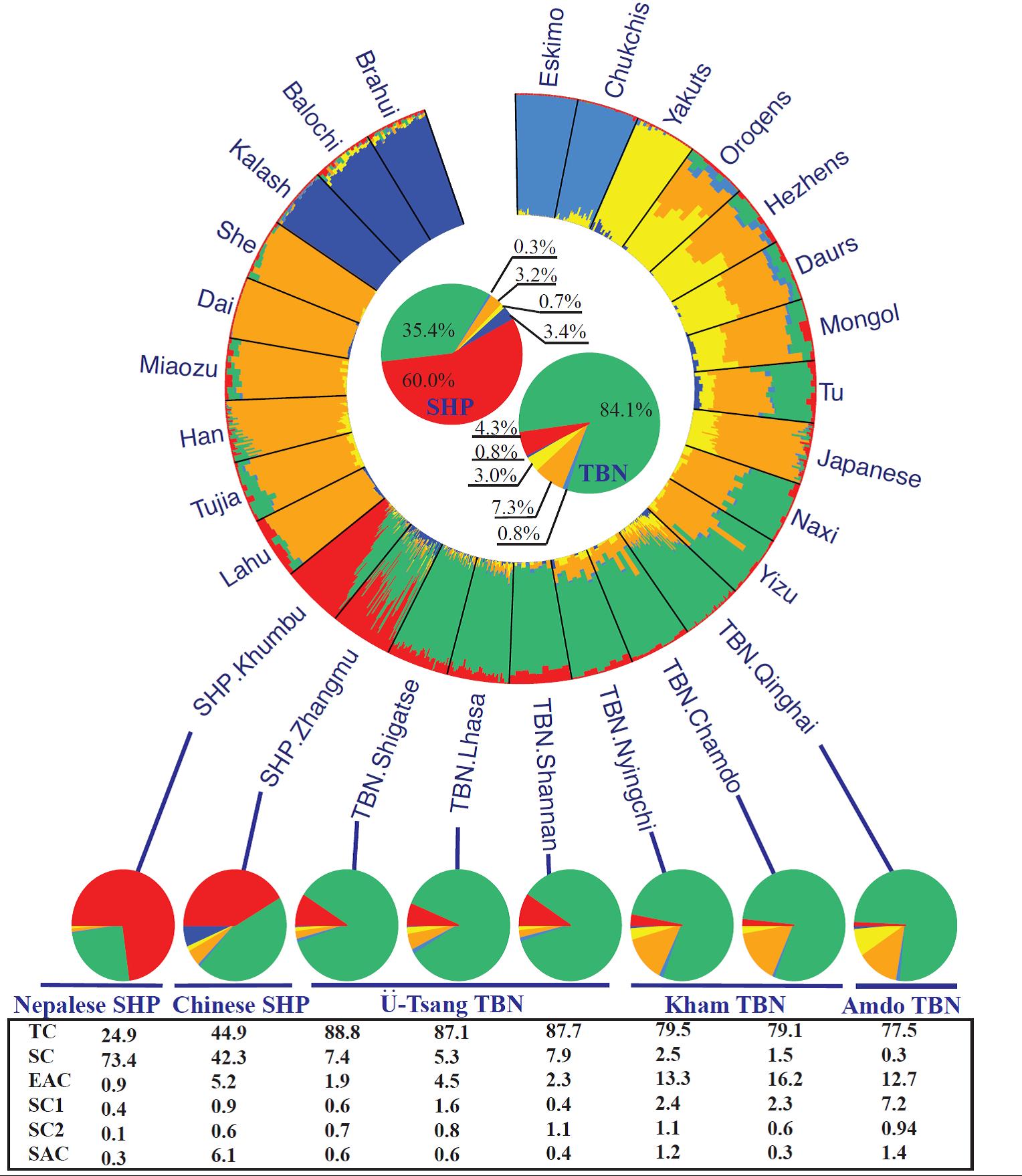
To resolve this complexity, Zhang et al. examined a wider geographic region, analyzing whole-genome sequences and genotyping data from Tibetans (from Tibet and Qinghai) and Sherpas (from Tibet-Zhangmu and Nepal-Khumbu). The sampling coverage from both of the populations in overlapping region was a promising step to rule out allele sharing via the isolation by distance model.
Although a substructure exists among both Tibetans and Sherpa, they are two genetically distinct populations. Sherpas have significant South Asian ancestry compared with Tibetans.
Interestingly, and in contrast with their geographical placement, Tibetan (Zhangmu) Sherpa had higher South Asian ancestry than Nepalese (Khumbu) Sherpa, which may be caused by lower amount of gene flow in the Nepalese (Khumbu) Sherpa, who live at a higher altitude. This population also had high consanguinity and a low effective population size – a further reflection of their geographic isolation.
In fact, the authors suggest that other components (e.g. East Asian and Siberian/Central Asian) in Sherpa genomes are likely caused by indirect gene flow from Tibetans.
It is noteworthy that this study inferred a strong bottleneck among Sherpa around 320-360 generations ago that is not found in the genetic history of Tibetans.
The authors estimate that Nepalese Sherpa diverged from Han Chinese 9,500-18,000 years ago, and from Tibetans 6,100-13,300 years ago. Meanwhile, Nepalese Sherpa and Tibetan Sherpa show a more recent divergence time of 1,200-7,800 years ago.
This study also identified novel Sherpa-specific genes (e.g. ALDH3A1, ANGPT1), which may contribute to high-altitude adaptation.
Gyaneshwer Chaubey
Latest posts by Gyaneshwer Chaubey (see all)
- Piecing together genetic histories of isolated populations - 21st June 2017
Very well written in layman’s language. Thanks.
Very illuminating article. Indian Government should do some serious work, otherwise we may not see any Parsi people in next 100 years.
The distinction of Tibetan and Sherpa is very interesting. I have seem many Sherps and indeed they are ‘king of mountains’.