Dr Hayley Bennett is a researcher from the parasite genomics group at the Wellcome Trust Sanger Institute. She is the lead author of an article published in Genome Biology which has revealed the genome of a rare tapeworm found living inside a patient’s brain. In this post she talks about new developments in genome sequencing that are managing to reveal an impressive amount of detail on potential drug targets for rare infections.
We have recently collaborated with pathologists, radiologists and clinicians looking at an exceptionally rare case of a tapeworm in a patient’s brain. The worm was removed by surgery and the material was used to find out more about a hitherto unsequenced order of tapeworms.
The case had baffled clinicians who tested for all the known diagnostic possibilities in the UK, until the worm was removed by surgical biopsy. One diagnosis that they had considered pre-surgery was infection with Taenia solium, a more common tapeworm that can encyst in the brain, however tests for this specific worm proved negative. Unlike in neurocysticercosis, where larval Taenia worms are contained within cysts, it appears that this worm had a free rein, travelling about 5 cm between the two hemispheres of the brain in a 4-year period.
Based on the physical characteristics of the worm, the infection was identified as sparganosis, which has not been observed in the UK before. It is thought that sparganosis can be caught by eating a raw amphibian or reptile that contains a motile larva form called a sparganum. It is so rare that only 300 cases had been reported prior to 2003, predominantly in China and South-East Asia.
-
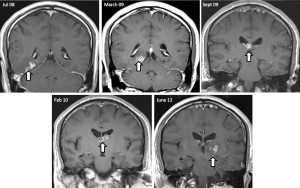
Brain MRI Scan showing migration of tapeworm over time. Image credit: Nagui Antoun (Click to enlarge)
Because of the scarcity of genetic information on different isolates, we suspect but cannot prove, that the infection was acquired during the patient’s travels in this region. We used a mitochondrial gene, cox1 known as the “barcode of life” to first identify the exact causative species as Spirometra erinaceieuropaei, distinguishing it from the proliferative Sparganum proliferum, which would have had a poorer outcome for the patient.
With the species established we asked what else we could learn from this case. Even though we only had a very small quantity of worm DNA from the biopsy sample, just under 50 nanograms, we were able to sequence and assemble a genome – a testament to the improvements in sequencing and bioinformatics methodology that have been made in recent years.
Using the genome we examined what types of genes were most abundant in Spirometra erinaceieuropaei, in comparison to other tapeworms, with the aim of working out what genes might be particularly important to this worm. As this worm was the first of its order to be sequenced, we expected it to be quite different. We found lots of gene families in Spirometra erinaceieuropaei with more gene copies than in other tapeworms. These families included several types of proteases, which may help the worm to digest different host tissues in order to invade or to feed. We also found lots of genes that are involved in modifying the proteins found within the worm, some of these may account for the complex changes in the worm and its ability to survive different environments throughout its lifecycle.
We were also able to look at the genes that represent potential drug targets identified in the genomes of other tapeworms, and found many that were also present in Spirometra erinaceieuropaei. As new treatments are developed to combat the more common tapeworms, it is our hope that the genome will serve as an ongoing reference for the research community. Researchers will be able to consult the genome and predict which drugs can be successfully repurposed for rarer infections.
There is still much to understand about this particular worm and tapeworms in general. The genome provides a great reference book for the organism, telling us what genes are present, but further research into when and how genes are used by tapeworms should give us even more insight into these complex parasites. Finally, our research highlights how astute pathology in combination with the latest genomics techniques can successfully push the boundaries of our current knowledge.
Latest posts by Guest (see all)
- Microbial forensics: It’s not just fingerprints that can be left behind - 12th May 2015
- How can ‘conservation genomics’ help the recovery of the most endangered species? - 12th December 2014
- Acetate helps hypoxic cancer cells get fat - 11th December 2014
Comments