Genome Medicine introduces a new series on Cancer epigenomics – the first articles include an editorial from Guest Editor Stephan Beck, a comment from Christoph Bock and a Q&A from Andrew Feinberg, as well as research from Christoph Plass and colleagues.
Recently, Cancer Research UK reported that 50% of people currently diagnosed with cancer will survive for at least 10 years. This compares to just 25% in the early 1970s, when the then US President Richard Nixon signed the National Cancer Act, initiating a ‘war on cancer’ and paving the way for major national and international funding. A cure for the different types of cancer is still elusive, but the achievements of the Human Genome Project and subsequent large-scale cancer-focused initiatives have raised hopes and expectations, sustained by the popular press.
 So it can be argued that the war against some cancer types is slowly being won in terms of extended survival, as a result of improved conventional treatments and new targeted therapies. Promising results with several new drugs were recently presented at the American Society of Clinical Oncology Annual Meeting in Chicago. However, the long-term prospects for many cancers after diagnosis remain dismal.
So it can be argued that the war against some cancer types is slowly being won in terms of extended survival, as a result of improved conventional treatments and new targeted therapies. Promising results with several new drugs were recently presented at the American Society of Clinical Oncology Annual Meeting in Chicago. However, the long-term prospects for many cancers after diagnosis remain dismal.
Next-generation sequencing and other omic approaches are now leading to a better understanding of specific cancer types, and these are already impacting clinical management and are heralding a new era of personalized precision oncology. While a cure for different cancer types remains an important but uncertain goal, efforts aimed at effective clinical management are now starting to emerge as the new holy grail of clinical cancer care.
The C2c approach
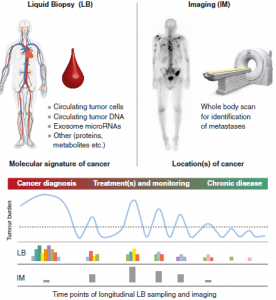 In his editorial, Stephan Beck introduces the C2c (cancer to chronic disease) approach, an initiative that will get underway in the coming months. It proposes that the new goal in oncology should be to turn cancer into a clinically manageable chronic disease, keeping the acute and aggressive forms of disease under sustained control throughout life after an initial diagnosis, similar to how diabetes is currently managed.
In his editorial, Stephan Beck introduces the C2c (cancer to chronic disease) approach, an initiative that will get underway in the coming months. It proposes that the new goal in oncology should be to turn cancer into a clinically manageable chronic disease, keeping the acute and aggressive forms of disease under sustained control throughout life after an initial diagnosis, similar to how diabetes is currently managed.
Beck uses the analogy of Dr. Faust’s pact with the devil, signed in blood, in his search for ultimate knowledge, as it is potentially through blood that the C2c approach could succeed. The approach involves multidimensional omic analyses of liquid biopsies (such as blood samples or other minimally invasive samples) in combination with state-of-the-art whole-body molecular imaging. Together, these allow longitudinal monitoring of patients, to ultimately enable targeted treatment of primary and secondary disease, and the prevention of subsequent relapses.
Cancer epigenomics
Our new series aims to highlight how advances driven by genome-wide and high-throughput technologies are allowing the detection of clinically relevant epigenetic alterations and the development of new approaches for detecting, monitoring and managing cancer. The series will feature research, alongside specially commissioned articles by leaders in the field.
International collaborative genomic studies involving high-throughput sequencing of different types of cancer have so far revealed complex patterns of mutations associated with initiation and progression of disease. However, gene expression and cellular function are also greatly impacted by the epigenome, and recent studies have revealed the importance of epigenetic alterations in cancer, including DNA methylation and histone modifications. The processes involved in generating these reversible marks are influenced by the environment, such as changes in cellular metabolism.
Drugs targeting regulators of epigenetic marks, such as DNA methyltransferase (DNMT) inhibitors and histone deacetylase (HDAC) inhibitors have shown efficacy in treating specific cancers. Several have been approved by the US FDA for treatment of specific hematologic cancers, including HDAC inhibitors vorinostat and romidepsin and DNMT inhibitors azacitidine and decitabine.
Other drugs that target regulators of epigenetic alterations are currently undergoing preclinical and clinical investigation for cancer therapy. However, further efforts are needed to understand the role of epigenetic alterations in cancer development and progression, before their potential as biomarkers and as targets for intervention can be fully realized.
Among the main challenges in the field is the integration of genomic and epigenomic information to understand their individual and combined roles in cancer. In a Comment, Christoph Bock argues that large-scale projects in the fields of cancer genomics and epigenomics need to work more closely together, and that integration of the resulting findings will allow better understanding of cancer biology and bring new translational opportunities. Bock also advocates the creation of an International Cancer Epigenome Consortium to facilitate more comprehensive characterization of cancer epigenomes.
DNA methylation signatures
DNA methylation is the best studied epigenetic alteration in cancer. In a Q&A, Andrew Feinberg provides some interesting insights into how DNA methylation contributes to cancer, highlighting the role of variability in DNA methylation, as well as his previously proposed epigenetic progenitor model. In this model, cancer arises in three steps, starting with epigenetic alterations of stem/progenitor cells in a tissue, followed by mutations in genes involved in cancer initiation, and then genetic and epigenetic instability that lead to increased tumor evolution and heterogeneity.
 Feinberg also highlights recent research that supports this model, including work by Teschendorff, Widschwendter and colleagues demonstrating that DNA methylation profiles in normal cells of origin of an epithelial cancer could be used to predict the risk of developing such a cancer.
Feinberg also highlights recent research that supports this model, including work by Teschendorff, Widschwendter and colleagues demonstrating that DNA methylation profiles in normal cells of origin of an epithelial cancer could be used to predict the risk of developing such a cancer.
Alongside better clinical management of cancer, prevention strategies are also a major goal. These have so far included extreme measures such as prophylactic surgery – Angelina Jolie’s double mastectomy one year ago raised public awareness of this as an option. A better understanding of the epigenetic processes involved in initiation and development of cancer, including DNA methylation, will be needed if such approaches are to be avoided in future cancer prevention strategies.
Given the importance of DNA methylation signatures, another major challenge is the development of useful preclinical models of cancer involving such signatures to test new approaches for intervention. In a new research paper, Christoph Plass and colleagues investigated DNA methylation patterns in a mouse model of acute myeloid leukemia (AML). They profiled genome-wide patterns of DNA methylation at three stages of disease progression (preleukemic, early and late leukemia) in their mouse model, and identified aberrantly methylated genes associated with early disease, which were also found in patients with AML. Their results indicate the utility of this model for studying DNA methylation signatures in patients with AML.
Looking ahead
Despite recent efforts to target epigenetic alterations, many hurdles remain, including the challenge of tumor heterogeneity. The latter may be resolved through single cell approaches to identify genomic and epigenomic alterations in circulating tumor cells via liquid biopsies, as part of Beck’s C2c approach.
Advances in cancer epigenomics, together with genomic advances in understanding cancer progression (the focus of this year’s Beyond the Genome conference, organized by Genome Medicine and Genome Biology) are expected to revolutionize clinical cancer management, and we look forward to highlighting these in our upcoming issues of Genome Medicine.
There is still time to submit research for the Cancer epigenomics series – the deadline for submissions is August 1st 2014. We look forward to receiving your feedback on the content, and on any additional topics that you would like to see featured.
Comments