Written by Dr Adele Murrell, University of Bath, UK
A long time ago, when we still used restriction enzymes to identify polymorphisms and the term “epigenetic” was used to dismiss quirky data that did not obey the central dogma of ‘DNA-makes-RNA-makes-protein’, we used to think of genes as linear sequences of bases in a sea of junk DNA. At bit more than a decade ago, just as the human genome sequencing project was nearing completion, the term “epigenetic” was confined to instances of stable mitotic inheritance of gene expression changes that could not be attributed to changes in nucleotide sequence. DNA methylation and histone modifications associate with gene expression changes and most epigenetic studies involved analysis of DNA methylation and possibly some histone acetylation. Nowadays, in order to be a card-carrying epigeneticist, one has to bear in mind that the linear nucleotide gene sequences are actually wrapped around nucleosomes to form chromatin that adopts open and closed conformations to regulate gene expression. Chromatin itself is further packaged into spatially segregated megabase-sized domains and sub-megabase-sized topological domains which may well form the architectural scaffold of chromosomes. Gene regulatory elements such as promoters and enhancers separated by long genomic distances can be brought closer by chromatin folding (looping) and the base of these loops are conceptualised as topological associated domains (TADs).
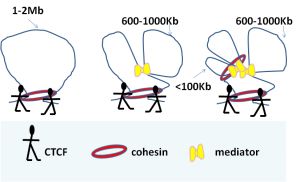
Initial studies of chromatin looping conformation focused on single loci. These studies identified a pivotal role for the 11 zinc finger CCCTC- binding factor, CTCF, in shaping chromatin loops and demonstrated that cohesin co-localised with CTCF to stabilise loops 1-5. Cohesin was also shown to interact with large promoter-enhancer complexes such as Mediator 6. In the last two years our understanding of chromatin organisation has been further increased by studies that have taken a genome-wide approach. It has been noted that architectural proteins CTCF and/or cohesin binding sites are enriched at boundaries between TADS 7; 8. However, CTCF and cohesin sites are also present within TADS and may therefore not be the only determinants of TAD structure.
In the June issue of Cell this year, Phillips-Cremins et al. identified that distinct combinations of CTCF, CTCF, cohesin, and Mediator work in a combinatorial manner to functionally organize chromatin in a cell-type-specific manner at the submegabase-length scale 9. This was done using a 5C-seq strategy7, to generate high-resolution maps of chromatin interactions across seven genomic regions in mouse embryonic stem cells (ESCs) and neural progenitor cells (NPCs). What makes this study remarkable is that the resolution is at 4kb fragment level, which is substantially higher than previous Hi-C techniques (average 40 kb resolution) and therefore revealed sub-topologically associated domains. The results showed that some TADs were invariant across different cell types, but many cell-specific differences in chromatin organisation were revealed within TADs at the sub-megabase level9. Interactions over short genomic distances (<100kb) were predominantly between promoters and enhancers and were bridged by cohesin and Mediator. At intermediate distances (<300) CTCF, cohesin and Mediator were often the bridging proteins, while at 600 – 1000kb Mediator complexes seem to hold loops together. CTCF alone or CTCF together with cohesin bridged interactions that were larger than 1MB. It will be interesting to see whether the paradigms for hierarchical differences in chromatin topology as observed between ESC and NCPs will be applicable in all cell types. My prediction is that it will. In fact, earlier this year Seitan et al found that depletion of cohesin in non-cycling thymocytes has no effect on the formation of Mb-compartmentalisation of the genome, but does affect specific promoter-enhancer interactions,10 which would fit in with the above observation of interactions at shorter genomic distances. Interestingly, reduction of interactions at the sub-megabase level increased or reinforced interactions within the larger domains suggesting that conformational changes in one region influence the conformation at a wider region10.
The frequency of occurrence of aberrant chromatin topography in disease is still unknown. Since chromatin organisation is a ubiquitous feature of all cells, mutations in genes that affect chromatin organisation are expected to be rare. Cohesin subunits and associated proteins are involved in a spectrum of developmental disorders known collectively as cohesinopathies. The most well-known cohesinopathy is Cornelia de Lange syndrome (CdLS). Although cohesin is best known for its role in holding sister chromatids together in mitosis, most CdLS patients show alterations in gene regulation rather than mitotic defects 11. The Bickmore lab has found that NIPBL, a protein responsible for loading the cohesin complex onto chromatin during S-phase, may have a role in regulating higher order chromatin structure, independent of CTCF and cohesin12. CdLS patients with mutations in the NIPBL gene show chromatin decompaction, which is visible by fluorescent in situ hybridisation12 . No mutations had been reported in CTCF until recently. Gregor et al have now discovered mutations in the CTCF gene in individuals with intellectual impairment and growth defects13. These mutations result in functional haplo-insuffiency of CTCF, and transcriptome data from affected patients identified several down-regulated genes. Consultation of ChIA-PET interaction data showed that the down-regulated genes in these patients could feasibly be due to changes in chromatin topology13.
Without even mentioning cancer or complex acquired disease where global epigenetic gene regulation is disrupted, it is clear that we are on the brink of understanding the contribution of aberrant chromatin conformation to disease formation. The next step is to develop easy user-friendly assays to identify aberrant chromatin conformation on a variety of patient tissue samples.
References:
1. Hadjur, S., Williams, L.M., Ryan, N.K., Cobb, B.S., Sexton, T., Fraser, P., Fisher, A.G., and Merkenschlager, M. (2009). Cohesins form chromosomal cis-interactions at the developmentally regulated IFNG locus. Nature 460, 410-413.
2. Mishiro, T., Ishihara, K., Hino, S., Tsutsumi, S., Aburatani, H., Shirahige, K., Kinoshita, Y., and Nakao, M. (2009). Architectural roles of multiple chromatin insulators at the human apolipoprotein gene cluster. Embo J 28, 1234-1245.
3. Degner, S.C., Verma-Gaur, J., Wong, T.P., Bossen, C., Iverson, G.M., Torkamani, A., Vettermann, C., Lin, Y.C., Ju, Z., Schulz, D., et al. (2011). CCCTC-binding factor (CTCF) and cohesin influence the genomic architecture of the Igh locus and antisense transcription in pro-B cells. Proc Natl Acad Sci U S A 108, 9566-9571.
4. Chien, R., Zeng, W., Ball, A.R., and Yokomori, K. (2011). Cohesin: a critical chromatin organizer in mammalian gene regulation. Biochem Cell Biol 89, 445-458.
5. Nativio, R., Wendt, K.S., Ito, Y., Huddleston, J.E., Uribe-Lewis, S., Woodfine, K., Krueger, C., Reik, W., Peters, J.M., and Murrell, A. (2009). Cohesin is required for higher-order chromatin conformation at the imprinted IGF2-H19 locus. PLoS Genet 5, e1000739.
6. Kagey, M.H., Newman, J.J., Bilodeau, S., Zhan, Y., Orlando, D.A., van Berkum, N.L., Ebmeier, C.C., Goossens, J., Rahl, P.B., Levine, S.S., et al. (2010). Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430-435.
7. Dixon, J.R., Selvaraj, S., Yue, F., Kim, A., Li, Y., Shen, Y., Hu, M., Liu, J.S., and Ren, B. (2012). Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376-380.
8. Nora, E.P., Lajoie, B.R., Schulz, E.G., Giorgetti, L., Okamoto, I., Servant, N., Piolot, T., van Berkum, N.L., Meisig, J., Sedat, J., et al. (2012). Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381-385.
9. Phillips-Cremins, J.E., Sauria, M.E., Sanyal, A., Gerasimova, T.I., Lajoie, B.R., Bell, J.S., Ong, C.T., Hookway, T.A., Guo, C., Sun, Y., et al. (2013). Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281-1295.
10. Seitan, V., Faure, A., Zhan, Y., McCord, R., Lajoie, B., Ing-Simmons, E., Lenhard, B., Giorgetti, L., Heard, E., Fisher, A., et al. (2013). Cohesin-based chromatin interactions enable regulated gene expression within pre-existing architectural compartments. Genome Res.
11. Castronovo, P., Gervasini, C., Cereda, A., Masciadri, M., Milani, D., Russo, S., Selicorni, A., and Larizza, L. (2009). Premature chromatid separation is not a useful diagnostic marker for Cornelia de Lange syndrome. Chromosome research : an international journal on the molecular, supramolecular and evolutionary aspects of chromosome biology 17, 763-771.
12. Nolen, L.D., Boyle, S., Ansari, M., Pritchard, E., and Bickmore, W.A. (2013). Regional chromatin decompaction in Cornelia de Lange syndrome associated with NIPBL disruption can be uncoupled from cohesin and CTCF. Hum Mol Genet.
13. Gregor, A., Oti, M., Kouwenhoven, E.N., Hoyer, J., Sticht, H., Ekici, A.B., Kjaergaard, S., Rauch, A., Stunnenberg, H.G., Uebe, S., et al. (2013). De novo mutations in the genome organizer CTCF cause intellectual disability. American journal of human genetics 93, 124-131.
Sam Rose
Latest posts by Sam Rose (see all)
- Raising funds for genetic diseases - 23rd September 2016
- The Epigenetics and Chromatin Clinic - 9th November 2015
- Resurrecting one of the oldest genetics journals - 23rd October 2015
Comments